TALEN-Based HvMPK3 Knock-Out Attenuates Proteome and Root Hair Phenotypic Responses to flg22 in Barley
- PMID: 33995462
- PMCID: PMC8117018
- DOI: 10.3389/fpls.2021.666229
TALEN-Based HvMPK3 Knock-Out Attenuates Proteome and Root Hair Phenotypic Responses to flg22 in Barley
Abstract
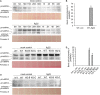
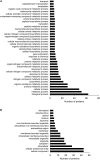
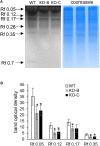
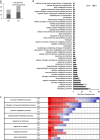
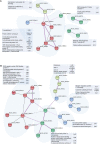
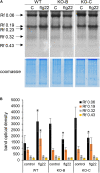
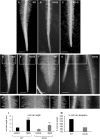
Mitogen activated protein kinases (MAPKs) integrate elicitor perception with both early and late responses associated with plant defense and innate immunity. Much of the existing knowledge on the role of plant MAPKs in defense mechanisms against microbes stems from extensive research in the model plant Arabidopsis thaliana. In the present study, we investigated the involvement of barley (Hordeum vulgare) MPK3 in response to flagellin peptide flg22, a well-known bacterial elicitor. Using differential proteomic analysis we show that TALEN-induced MPK3 knock-out lines of barley (HvMPK3 KO) exhibit constitutive downregulation of defense related proteins such as PR proteins belonging to thaumatin family and chitinases. Further analyses showed that the same protein families were less prone to flg22 elicitation in HvMPK3 KO plants compared to wild types. These results were supported and validated by chitinase activity analyses and immunoblotting for HSP70. In addition, differential proteomes correlated with root hair phenotypes and suggested tolerance of HvMPK3 KO lines to flg22. In conclusion, our study points to the specific role of HvMPK3 in molecular and root hair phenotypic responses of barley to flg22.
Keywords: HvMPK3; PR proteins; TALEN; barley; chitinases; flagellin; proteomics; root hairs.
Copyright © 2021 Takáč, Křenek, Komis, Vadovič, Ovečka, Ohnoutková, Pechan, Kašpárek, Tichá, Basheer, Arick and Šamaj.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

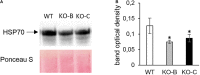
Similar articles
-
Knockout of MITOGEN-ACTIVATED PROTEIN KINASE 3 causes barley root resistance against Fusarium graminearum.Plant Physiol. 2022 Nov 28;190(4):2847-2867. doi: 10.1093/plphys/kiac389. Plant Physiol. 2022. PMID: 35993881 Free PMC article.
-
Functional analysis of Arabidopsis immune-related MAPKs uncovers a role for MPK3 as negative regulator of inducible defences.Genome Biol. 2014 Jun 30;15(6):R87. doi: 10.1186/gb-2014-15-6-r87. Genome Biol. 2014. PMID: 24980080 Free PMC article.
-
Activation of the Arabidopsis thaliana mitogen-activated protein kinase MPK11 by the flagellin-derived elicitor peptide, flg22.Mol Plant Microbe Interact. 2012 Apr;25(4):471-80. doi: 10.1094/MPMI-11-11-0281. Mol Plant Microbe Interact. 2012. PMID: 22204645
-
Comparative analysis of stress-induced calcium signals in the crop species barley and the model plant Arabidopsis thaliana.BMC Plant Biol. 2022 Sep 17;22(1):447. doi: 10.1186/s12870-022-03820-5. BMC Plant Biol. 2022. PMID: 36114461 Free PMC article.
-
Magical mystery tour: MLO proteins in plant immunity and beyond.New Phytol. 2014 Oct;204(2):273-81. doi: 10.1111/nph.12889. New Phytol. 2014. PMID: 25453131 Review.
Cited by
-
Methyl viologen-induced changes in the Arabidopsis proteome implicate PATELLIN 4 in oxidative stress responses.J Exp Bot. 2024 Jan 1;75(1):405-421. doi: 10.1093/jxb/erad363. J Exp Bot. 2024. PMID: 37728561 Free PMC article.
-
Knockout of MITOGEN-ACTIVATED PROTEIN KINASE 3 causes barley root resistance against Fusarium graminearum.Plant Physiol. 2022 Nov 28;190(4):2847-2867. doi: 10.1093/plphys/kiac389. Plant Physiol. 2022. PMID: 35993881 Free PMC article.
-
CRISPR/Cas9-Induced Loss-of-Function Mutation in the Barley Mitogen-Activated Protein Kinase 6 Gene Causes Abnormal Embryo Development Leading to Severely Reduced Grain Germination and Seedling Shootless Phenotype.Front Plant Sci. 2021 Jul 30;12:670302. doi: 10.3389/fpls.2021.670302. eCollection 2021. Front Plant Sci. 2021. PMID: 34394137 Free PMC article.
-
Getting defensive: Kinase MPK3 offers a target for breeding Fusarium-resistant barley.Plant Physiol. 2023 Jan 2;191(1):26-28. doi: 10.1093/plphys/kiac473. Plant Physiol. 2023. PMID: 36219040 Free PMC article. No abstract available.
-
A remodeled RNA polymerase II complex catalyzing viroid RNA-templated transcription.PLoS Pathog. 2022 Sep 19;18(9):e1010850. doi: 10.1371/journal.ppat.1010850. eCollection 2022 Sep. PLoS Pathog. 2022. PMID: 36121876 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

