Genome-wide screening of circadian and non-circadian impact of Neat1 genetic deletion
- PMID: 33995907
- PMCID: PMC8085668
- DOI: 10.1016/j.csbj.2021.04.022
Genome-wide screening of circadian and non-circadian impact of Neat1 genetic deletion
Abstract
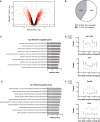
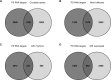
The functions of the long non-coding RNA, Nuclear enriched abundant transcript 1 (Neat1), are poorly understood. Neat1 is required for the formation of paraspeckles, but its respective paraspeckle-dependent or independent functions are unknown. Several studies including ours reported that Neat1 is involved in the regulation of circadian rhythms. We characterized the impact of Neat1 genetic deletion in a rat pituitary cell line. The mRNAs whose circadian expression pattern or expression level is regulated by Neat1 were identified after high-throughput RNA sequencing of the circadian transcriptome of wild-type cells compared to cells in which Neat1 was deleted by CRISPR/Cas9. The numerous RNAs affected by Neat1 deletion were found to be circadian or non-circadian, targets or non-targets of paraspeckles, and to be associated with many key biological processes showing that Neat1, in interaction with the circadian system or independently, could play crucial roles in key physiological functions through diverse mechanisms.
Keywords: CRISPR/Cas9 edition; Circadian gene expression; Long non-coding RNA Neat1; Paraspeckles; RNA-sequencing.
© 2021 The Author(s).
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



References
-
- Riva P., Ratti A., Venturin M. The long non-coding RNAs in neurodegenerative diseases: novel mechanisms of pathogenesis. Curr Alzheimer Res. 2016;13:1219–1231. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
