Molecular mechanism of secreted amyloid-β precursor protein in binding and modulating GABABR1a
- PMID: 33996007
- PMCID: PMC8098695
- DOI: 10.1039/d0sc06946a
Molecular mechanism of secreted amyloid-β precursor protein in binding and modulating GABABR1a
Abstract
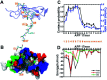
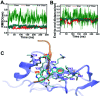
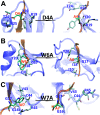
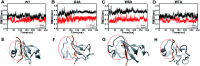
A recent phenomenal study discovered that the extension domain of secreted amyloid-β precursor protein (sAPP) can bind to the intrinsically disordered sushi 1 domain of the γ-aminobutyric acid type B receptor subunit 1a (GABABR1a) and modulate its synaptic transmission. The work provided an important structural foundation for the modulation of GABABR1a; however, the detailed molecular interaction mechanism, crucial for future drug design, remains elusive. Here, we further investigated the dynamical interactions between sAPP peptides and the natively unstructured sushi 1 domain using all-atom molecular dynamics simulations, for both the 17-residue sAPP peptide (APP 17-mer) and its minimally active 9 residue segment (APP 9-mer). We then explored mutations of the APP 9-mer with rigorous free energy perturbation (FEP) calculations. Our in silico mutagenesis studies revealed key residues (D4, W6, and W7) responsible for the binding with the sushi 1 domain. More importantly, one double mutation based on different vertebrate APP sequences from evolution exhibited a stronger binding (ΔΔG = -1.91 ± 0.66 kcal mol-1), indicating a potentially enhanced GABABR1a modulator. These large-scale simulations may provide new insights into the binding mechanism between sAPP and the sushi 1 domain, which could open new avenues in the development of future GABABR1a-specific therapeutics.
This journal is © The Royal Society of Chemistry.
Conflict of interest statement
There are no conflicts to declare.
Figures
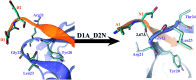
Similar articles
-
Secreted amyloid-β precursor protein functions as a GABABR1a ligand to modulate synaptic transmission.Science. 2019 Jan 11;363(6423):eaao4827. doi: 10.1126/science.aao4827. Science. 2019. PMID: 30630900 Free PMC article.
-
Identification of the probable structure of the sAPPα-GABABR1a complex and theoretical solutions for such cases.Phys Chem Chem Phys. 2022 May 25;24(20):12267-12280. doi: 10.1039/d2cp00569g. Phys Chem Chem Phys. 2022. PMID: 35543350
-
Toxicity Mechanism of Aβ42 Oligomer in the Binding between the GABABR1a sushi1 Domain and Amyloid Precursor Protein 9mer: A Mechanism like Substitution Reaction.ACS Chem Neurosci. 2022 Jul 6;13(13):2048-2059. doi: 10.1021/acschemneuro.2c00311. Epub 2022 Jun 23. ACS Chem Neurosci. 2022. PMID: 35737468
-
Biologically active domain of the secreted form of the amyloid beta/A4 protein precursor.Ann N Y Acad Sci. 1993 Sep 24;695:149-57. doi: 10.1111/j.1749-6632.1993.tb23044.x. Ann N Y Acad Sci. 1993. PMID: 8239274 Review.
-
Cellular actions of beta-amyloid precursor protein and its soluble and fibrillogenic derivatives.Physiol Rev. 1997 Oct;77(4):1081-132. doi: 10.1152/physrev.1997.77.4.1081. Physiol Rev. 1997. PMID: 9354812 Review.
Cited by
-
Mutagenesis-based optimal design of plant peptide phytosulfokine for enhanced biological activity.Comput Struct Biotechnol J. 2025 Mar 24;27:1296-1304. doi: 10.1016/j.csbj.2025.03.029. eCollection 2025. Comput Struct Biotechnol J. 2025. PMID: 40230409 Free PMC article.
-
Probing the fluorination effect on the self-assembly characteristics, in vivo fate and antitumor efficacy of paclitaxel prodrug nanoassemblies.Theranostics. 2021 Jul 6;11(16):7896-7910. doi: 10.7150/thno.61337. eCollection 2021. Theranostics. 2021. PMID: 34335971 Free PMC article.
-
Evolution of Thyroglobulin Loop Kinetics in EpCAM.Life (Basel). 2021 Sep 3;11(9):915. doi: 10.3390/life11090915. Life (Basel). 2021. PMID: 34575064 Free PMC article.
-
Soluble amyloid-β precursor peptide does not regulate GABAB receptor activity.Elife. 2023 Jan 23;12:e82082. doi: 10.7554/eLife.82082. Elife. 2023. PMID: 36688536 Free PMC article.
References
-
- Rice H. C. de Malmazet D. Schreurs A. Frere S. Van Molle I. Volkov A. N. Creemers E. Vertkin I. Nys J. Ranaivoson F. M. Comoletti D. Savas J. N. Remaut H. Balschun D. Wierda K. D. Slutsky I. Farrow K. De Strooper B. de Wit J. Secreted amyloid-beta precursor protein functions as a GABA(B)R1a ligand to modulate synaptic transmission. Science. 2019;363(6423):143. - PMC - PubMed
-
- Golde T. E. Estus S. Younkin L. H. Selkoe D. J. Younkin S. G. Processing of the amyloid protein-precursor to potentially amyloidogenic derivatives. Science. 1992;255(5045):728–730. - PubMed
-
- Kang J. Lemaire H. G. Unterbeck A. Salbaum J. M. Masters C. L. Grzeschik K. H. Multhaup G. Beyreuther K. Mullerhill B. The precursor of Alzheimer's-disease amyloid-a4 protein resembles a cell-surface receptor. Nature. 1987;325(6106):733–736. - PubMed
-
- Willem M. Tahirovic S. Busche M. A. Ovsepian S. V. Chafai M. Kootar S. Hornburg D. Evans L. D. B. Moore S. Daria A. Hampel H. Mueller V. Giudici C. Nuscher B. Wenninger-Weinzierl A. Kremmer E. Heneka M. T. Thal D. R. Giedraitis V. Lannfelt L. Mueller U. Livesey F. J. Meissner F. Herms J. Konnerth A. Marie H. Haass C. eta-Secretase processing of APP inhibits neuronal activity in the hippocampus. Nature. 2015;526(7573):443–+. - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

