Abundance Imparts Evolutionary Constraints of Similar Magnitude on the Buried, Surface, and Disordered Regions of Proteins
- PMID: 33996892
- PMCID: PMC8119896
- DOI: 10.3389/fmolb.2021.626729
Abundance Imparts Evolutionary Constraints of Similar Magnitude on the Buried, Surface, and Disordered Regions of Proteins
Abstract
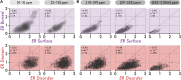
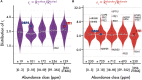
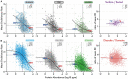
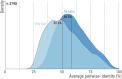
An understanding of the forces shaping protein conservation is key, both for the fundamental knowledge it represents and to allow for optimal use of evolutionary information in practical applications. Sequence conservation is typically examined at one of two levels. The first is a residue-level, where intra-protein differences are analyzed and the second is a protein-level, where inter-protein differences are studied. At a residue level, we know that solvent-accessibility is a prime determinant of conservation. By inverting this logic, we inferred that disordered regions are slightly more solvent-accessible on average than the most exposed surface residues in domains. By integrating abundance information with evolutionary data within and across proteins, we confirmed a previously reported strong surface-core association in the evolution of structured regions, but we found a comparatively weak association between disordered and structured regions. The facts that disordered and structured regions experience different structural constraints and evolve independently provide a unique setup to examine an outstanding question: why is a protein's abundance the main determinant of its sequence conservation? Indeed, any structural or biophysical property linked to the abundance-conservation relationship should increase the relative conservation of regions concerned with that property (e.g., disordered residues with mis-interactions, domain residues with misfolding). Surprisingly, however, we found the conservation of disordered and structured regions to increase in equal proportion with abundance. This observation implies that either abundance-related constraints are structure-independent, or multiple constraints apply to different regions and perfectly balance each other.
Keywords: contact number; intrinsic disorder; misfolding; misinteraction; protein abundance; protein evolution; protein structure; yeast proteome.
Copyright © 2021 Dubreuil and Levy.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Deciphering the cause of evolutionary variance within intrinsically disordered regions in human proteins.J Biomol Struct Dyn. 2017 Feb;35(2):233-249. doi: 10.1080/07391102.2016.1143877. Epub 2016 May 4. J Biomol Struct Dyn. 2017. PMID: 26790343
-
Comparable contributions of structural-functional constraints and expression level to the rate of protein sequence evolution.Biol Direct. 2008 Oct 7;3:40. doi: 10.1186/1745-6150-3-40. Biol Direct. 2008. PMID: 18840284 Free PMC article.
-
Protein Abundance Biases the Amino Acid Composition of Disordered Regions to Minimize Non-functional Interactions.J Mol Biol. 2019 Dec 6;431(24):4978-4992. doi: 10.1016/j.jmb.2019.08.008. Epub 2019 Aug 20. J Mol Biol. 2019. PMID: 31442477 Free PMC article.
-
Ensembles from Ordered and Disordered Proteins Reveal Similar Structural Constraints during Evolution.J Mol Biol. 2019 Mar 15;431(6):1298-1307. doi: 10.1016/j.jmb.2019.01.031. Epub 2019 Feb 5. J Mol Biol. 2019. PMID: 30731089
-
Correlation between protein abundance and sequence conservation: what do recent experiments say?Curr Opin Genet Dev. 2022 Dec;77:101984. doi: 10.1016/j.gde.2022.101984. Epub 2022 Sep 23. Curr Opin Genet Dev. 2022. PMID: 36162152 Review.
Cited by
-
A Conserved Core Region of the Scaffold NEMO is Essential for Signal-induced Conformational Change and Liquid-liquid Phase Separation.bioRxiv [Preprint]. 2023 May 25:2023.05.25.542299. doi: 10.1101/2023.05.25.542299. bioRxiv. 2023. Update in: J Biol Chem. 2023 Dec;299(12):105396. doi: 10.1016/j.jbc.2023.105396. PMID: 37292615 Free PMC article. Updated. Preprint.
-
A conserved core region of the scaffold NEMO is essential for signal-induced conformational change and liquid-liquid phase separation.J Biol Chem. 2023 Dec;299(12):105396. doi: 10.1016/j.jbc.2023.105396. Epub 2023 Oct 27. J Biol Chem. 2023. PMID: 37890781 Free PMC article.
-
Substitution Models of Protein Evolution with Selection on Enzymatic Activity.Mol Biol Evol. 2024 Feb 1;41(2):msae026. doi: 10.1093/molbev/msae026. Mol Biol Evol. 2024. PMID: 38314876 Free PMC article.
References
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

