LINC02257, an Enhancer RNA of Prognostic Value in Colon Adenocarcinoma, Correlates With Multi-Omics Immunotherapy-Related Analysis in 33 Cancers
- PMID: 33996902
- PMCID: PMC8121256
- DOI: 10.3389/fmolb.2021.646786
LINC02257, an Enhancer RNA of Prognostic Value in Colon Adenocarcinoma, Correlates With Multi-Omics Immunotherapy-Related Analysis in 33 Cancers
Abstract
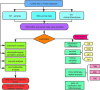
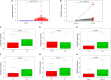
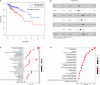
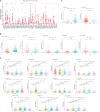
Accumulated evidence supports that long non-coding RNAs (lncRNAs) are involved significantly in the development of human cancers. Enhancer RNAs (eRNAs), a subtype of lncRNAs, have recently attracted much attention about their roles in carcinogenesis. Colon adenocarcinoma is one of the most commonly diagnosed tumors with unfavorable prognosis. It highlights the great significance of screening and identifying novel biomarkers. More importantly, it remains to be elucidated with respect to the function of eRNAs in colon adenocarcinoma, as is in pan-cancers. The expression of LINC02257 was determined based on the data obtained from The Cancer Genome Atlas (TCGA). Further evaluation was performed on the basis of the following analyses: clinicopathology and survival analysis, gene ontology (GO) terms, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, as well as multi-omics immunotherapy-related analysis and co-expression analysis. The statistical analysis was conducted in R software, and immune cell infiltration of LINC02257 expression in cancers was investigated by using the CIBERSORT algorithm. By large-scale data mining, our study highlighted that a total of 39 eRNA genes were associated with colon adenocarcinoma prognosis, among which 25 eRNAs showed significant associations with their predicted target genes. LINC02257 was identified as the most significant survival-associated eRNA, with DUSP10 as its target gene. Besides, the high expression of LINC02257 in colon adenocarcinoma was more vulnerable to unfavorable prognosis and correlated with various clinical characteristics. GO and KEGG analyses revealed that LINC02257 was closely correlated with extracellular matrix organization via the PI3K-Akt signaling pathway. Besides, LINC02257 expression correlated with a multi-omics analysis of 33 cancer types, such as survival analysis [overall survival (OS), disease-specific survival (DSS), disease-free interval (DFI), and progression-free interval (PFI)] and immunotherapy-related analysis [tumor microenvironment (TME), tumor mutational burden (TMB), and microsatellite instability (MSI)]. Finally, we investigated the co-expression genes of LINC02257 and its potential signaling pathways across different cancer types. LINC02257 is screened and can function as an independent prognostic biomarker through the PI3K-Akt signaling pathway for colon adenocarcinoma. Simultaneously, LINC02257 may be a multifaceted and significant immunotherapy-related eRNA in different cancers.
Keywords: LINC02257; bioinformatic analysis; enhancer; immune-related multi-omics analysis; pan-cancers.
Copyright © 2021 Xiao, Liu, Yi and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
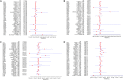
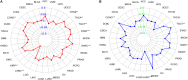
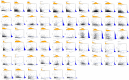
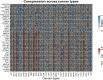
Figures

Similar articles
-
Expression characteristics of long non-coding RNA in colon adenocarcinoma and its potential value for judging the survival and prognosis of patients: bioinformatics analysis based on The Cancer Genome Atlas database.J Gastrointest Oncol. 2022 Jun;13(3):1178-1187. doi: 10.21037/jgo-22-384. J Gastrointest Oncol. 2022. PMID: 35837189 Free PMC article.
-
Identification of SHCBP1 as a potential biomarker involving diagnosis, prognosis, and tumor immune microenvironment across multiple cancers.Comput Struct Biotechnol J. 2022 Jun 18;20:3106-3119. doi: 10.1016/j.csbj.2022.06.039. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 35782736 Free PMC article.
-
Comprehensive characterization of functional eRNAs in lung adenocarcinoma reveals novel regulators and a prognosis-related molecular subtype.Theranostics. 2020 Sep 14;10(24):11264-11277. doi: 10.7150/thno.47039. eCollection 2020. Theranostics. 2020. PMID: 33042282 Free PMC article.
-
Pan-cancer analysis of the prognostic and immunological role of GJB2: a potential target for survival and immunotherapy.Front Oncol. 2023 Jun 23;13:1110207. doi: 10.3389/fonc.2023.1110207. eCollection 2023. Front Oncol. 2023. PMID: 37427102 Free PMC article. Review.
-
Emerging Role of Enhancer RNAs as Potential Diagnostic and Prognostic Biomarkers in Cancer.Noncoding RNA. 2022 Oct 1;8(5):66. doi: 10.3390/ncrna8050066. Noncoding RNA. 2022. PMID: 36287118 Free PMC article. Review.
Cited by
-
Enhancer RNAs (eRNAs) in Cancer: The Jacks of All Trades.Cancers (Basel). 2022 Apr 14;14(8):1978. doi: 10.3390/cancers14081978. Cancers (Basel). 2022. PMID: 35454885 Free PMC article. Review.
-
Expression characteristics of long non-coding RNA in colon adenocarcinoma and its potential value for judging the survival and prognosis of patients: bioinformatics analysis based on The Cancer Genome Atlas database.J Gastrointest Oncol. 2022 Jun;13(3):1178-1187. doi: 10.21037/jgo-22-384. J Gastrointest Oncol. 2022. PMID: 35837189 Free PMC article.
-
eRNA-IDO: A One-stop Platform for Identification, Interactome Discovery, and Functional Annotation of Enhancer RNAs.Genomics Proteomics Bioinformatics. 2024 Oct 15;22(4):qzae059. doi: 10.1093/gpbjnl/qzae059. Genomics Proteomics Bioinformatics. 2024. PMID: 39178387 Free PMC article.
-
A Hypoxia-Related lncRNA Signature Correlates with Survival and Tumor Microenvironment in Colorectal Cancer.J Immunol Res. 2022 Jul 8;2022:9935705. doi: 10.1155/2022/9935705. eCollection 2022. J Immunol Res. 2022. PMID: 35846431 Free PMC article.
-
Identification of Enhancer RNA CDK6-AS1 as a Potential Novel Prognostic Biomarker in Gastric Cancer.Front Genet. 2022 Apr 29;13:854211. doi: 10.3389/fgene.2022.854211. eCollection 2022. Front Genet. 2022. PMID: 35571043 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

