Structural Aspects of Auxin Signaling
- PMID: 34001533
- PMCID: PMC8725629
- DOI: 10.1101/cshperspect.a039883
Structural Aspects of Auxin Signaling
Abstract
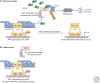
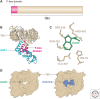
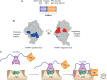
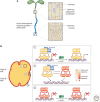
Auxin signaling regulates growth and developmental processes in plants. The core of nuclear auxin signaling relies on just three components: TIR1/AFBs, Aux/IAAs, and ARFs. Each component is itself made up of several domains, all of which contribute to the regulation of auxin signaling. Studies of the structural aspects of these three core signaling components have deepened our understanding of auxin signaling dynamics and regulation. In addition to the structured domains of these components, intrinsically disordered regions within the proteins also impact auxin signaling outcomes. New research is beginning to uncover the role intrinsic disorder plays in auxin-regulated degradation and subcellular localization. Structured and intrinsically disordered domains affect auxin perception, protein degradation dynamics, and DNA binding. Taken together, subtle differences within the domains and motifs of each class of auxin signaling component affect signaling outcomes and specificity.
Copyright © 2022 Cold Spring Harbor Laboratory Press; all rights reserved.
Figures

References
-
- Boer DR, Freire-Rios A, van den Berg WA, Saaki T, Manfield IW, Kepinski S, López-Vidrieo I, Franco-Zorrilla JM, de Vries SC, Solano R, et al. 2014. Structural basis for DNA binding specificity by the auxin-dependent ARF transcription factors. Cell 156: 577–589. 10.1016/j.cell.2013.12.027 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
