Spatio-temporal landscape of mouse epididymal cells and specific mitochondria-rich segments defined by large-scale single-cell RNA-seq
- PMID: 34001862
- PMCID: PMC8129088
- DOI: 10.1038/s41421-021-00260-7
Spatio-temporal landscape of mouse epididymal cells and specific mitochondria-rich segments defined by large-scale single-cell RNA-seq
Abstract
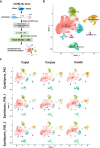
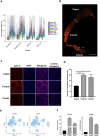
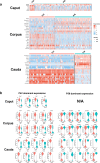
Spermatozoa acquire their fertilizing ability and forward motility during epididymal transit, suggesting the importance of the epididymis. Although the cell atlas of the epididymis was reported recently, the heterogeneity of the cells and the gene expression profile in the epididymal tube are still largely unknown. Considering single-cell RNA sequencing results, we thoroughly studied the cell composition, spatio-temporal differences in differentially expressed genes (DEGs) in epididymal segments and mitochondria throughout the epididymis with sufficient cell numbers. In total, 40,623 cells were detected and further clustered into 8 identified cell populations. Focused analyses revealed the subpopulations of principal cells, basal cells, clear/narrow cells, and halo/T cells. Notably, two subtypes of principal cells, the Prc7 and Prc8 subpopulations were enriched as stereocilia-like cells according to GO analysis. Further analysis demonstrated the spatially specific pattern of the DEGs in each cell cluster. Unexpectedly, the abundance of mitochondria and mitochondrial transcription (MT) was found to be higher in the corpus and cauda epididymis than in the caput epididymis by scRNA-seq, immunostaining, and qPCR validation. In addition, the spatio-temporal profile of the DEGs from the P42 and P56 epididymis, including transiting spermatozoa, was depicted. Overall, our study presented the single-cell transcriptome atlas of the mouse epididymis and revealed the novel distribution pattern of mitochondria and key genes that may be linked to sperm functionalities in the first wave and subsequent wave of sperm, providing a roadmap to be emulated in efforts to achieve sperm maturation regulation in the epididymis.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Region-specific gene expression in the epididymis of Yak.Theriogenology. 2019 Nov;139:132-146. doi: 10.1016/j.theriogenology.2019.08.006. Epub 2019 Aug 5. Theriogenology. 2019. PMID: 31404823
-
Distribution characteristics of mitochondria-rich segments in the epididymis.Am J Clin Exp Immunol. 2023 Aug 20;12(4):72-73. eCollection 2023. Am J Clin Exp Immunol. 2023. PMID: 37736076 Free PMC article.
-
Junctional adhesion molecule A: expression in the murine epididymal tract and accessory organs and acquisition by maturing sperm.Mol Hum Reprod. 2017 Feb 10;23(2):132-140. doi: 10.1093/molehr/gaw082. Mol Hum Reprod. 2017. PMID: 28062807 Free PMC article.
-
Role of exosomes in sperm maturation during the transit along the male reproductive tract.Blood Cells Mol Dis. 2005 Jul-Aug;35(1):1-10. doi: 10.1016/j.bcmd.2005.03.005. Blood Cells Mol Dis. 2005. PMID: 15893944 Review.
-
The current perspective on genetic and epigenetic factors in sperm maturation in the epididymis.Andrologia. 2021 Apr;53(3):e13989. doi: 10.1111/and.13989. Epub 2021 Jan 25. Andrologia. 2021. PMID: 33491190 Review.
Cited by
-
The Male Reproductive Toxicity Caused by 2-Naphthylamine Was Related to Testicular Immunity Disorders.Toxics. 2024 May 7;12(5):342. doi: 10.3390/toxics12050342. Toxics. 2024. PMID: 38787121 Free PMC article.
-
Molecular Pathways Implicated in the Differentiation and Function of Epididymal Basal Cells.Adv Exp Med Biol. 2025;1469:89-113. doi: 10.1007/978-3-031-82990-1_5. Adv Exp Med Biol. 2025. PMID: 40301254 Review.
-
Loss of ndrg2 Function Is Involved in Notch Activation in Neuromast Hair Cell Regeneration in Zebrafish.Mol Neurobiol. 2023 Jun;60(6):3100-3112. doi: 10.1007/s12035-023-03262-6. Epub 2023 Feb 17. Mol Neurobiol. 2023. PMID: 36800156
-
Single-cell transcriptome characteristics of testicular terminal epithelium lineages during aging in the Drosophila.Aging Cell. 2024 Mar;23(3):e14057. doi: 10.1111/acel.14057. Epub 2023 Dec 3. Aging Cell. 2024. PMID: 38044573 Free PMC article.
-
Calcium Homeostasis in the Epididymal Microenvironment: Is Extracellular Calcium a Cofactor for Matrix Gla Protein-Dependent Scavenging Regulated by Vitamins.Front Cell Dev Biol. 2022 Feb 17;10:827940. doi: 10.3389/fcell.2022.827940. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35252193 Free PMC article. Review.
References
-
- Turner TT. On the epididymis and its role in the development of the fertile ejaculate. J. Androl. 1995;16:292–298. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

