Transcriptome-wide association analysis of brain structures yields insights into pleiotropy with complex neuropsychiatric traits
- PMID: 34001886
- PMCID: PMC8128893
- DOI: 10.1038/s41467-021-23130-y
Transcriptome-wide association analysis of brain structures yields insights into pleiotropy with complex neuropsychiatric traits
Abstract
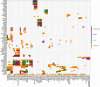
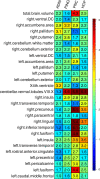
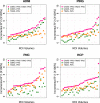
Structural variations of the human brain are heritable and highly polygenic traits, with hundreds of associated genes identified in recent genome-wide association studies (GWAS). Transcriptome-wide association studies (TWAS) can both prioritize these GWAS findings and also identify additional gene-trait associations. Here we perform cross-tissue TWAS analysis of 211 structural neuroimaging and discover 278 associated genes exceeding Bonferroni significance threshold of 1.04 × 10-8. The TWAS-significant genes for brain structures have been linked to a wide range of complex traits in different domains. Through TWAS gene-based polygenic risk scores (PRS) prediction, we find that TWAS PRS gains substantial power in association analysis compared to conventional variant-based GWAS PRS, and up to 6.97% of phenotypic variance (p-value = 7.56 × 10-31) can be explained in independent testing data sets. In conclusion, our study illustrates that TWAS can be a powerful supplement to traditional GWAS in imaging genetics studies for gene discovery-validation, genetic co-architecture analysis, and polygenic risk prediction.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Power analysis of transcriptome-wide association study: Implications for practical protocol choice.PLoS Genet. 2021 Feb 26;17(2):e1009405. doi: 10.1371/journal.pgen.1009405. eCollection 2021 Feb. PLoS Genet. 2021. PMID: 33635859 Free PMC article.
-
Bayesian genome-wide TWAS with reference transcriptomic data of brain and blood tissues identified 141 risk genes for Alzheimer's disease dementia.Alzheimers Res Ther. 2024 Jun 1;16(1):120. doi: 10.1186/s13195-024-01488-7. Alzheimers Res Ther. 2024. PMID: 38824563 Free PMC article.
-
Influence of tissue context on gene prioritization for predicted transcriptome-wide association studies.Pac Symp Biocomput. 2019;24:296-307. Pac Symp Biocomput. 2019. PMID: 30864331 Free PMC article. Clinical Trial.
-
Opportunities and challenges for transcriptome-wide association studies.Nat Genet. 2019 Apr;51(4):592-599. doi: 10.1038/s41588-019-0385-z. Epub 2019 Mar 29. Nat Genet. 2019. PMID: 30926968 Free PMC article. Review.
-
Polygenic risk score: use in migraine research.J Headache Pain. 2018 Apr 5;19(1):29. doi: 10.1186/s10194-018-0856-0. J Headache Pain. 2018. PMID: 29623444 Free PMC article. Review.
Cited by
-
Integration of estimated regional gene expression with neuroimaging and clinical phenotypes at biobank scale.PLoS Biol. 2024 Sep 13;22(9):e3002782. doi: 10.1371/journal.pbio.3002782. eCollection 2024 Sep. PLoS Biol. 2024. PMID: 39269986 Free PMC article.
-
Understanding Regulatory Mechanisms of Brain Function and Disease through 3D Genome Organization.Genes (Basel). 2022 Mar 25;13(4):586. doi: 10.3390/genes13040586. Genes (Basel). 2022. PMID: 35456393 Free PMC article. Review.
-
The landscape of plasma proteomic links to human organ imaging.medRxiv [Preprint]. 2025 Jan 15:2025.01.14.25320532. doi: 10.1101/2025.01.14.25320532. medRxiv. 2025. PMID: 39867388 Free PMC article. Preprint.
-
The eQTL colocalization and transcriptome-wide association study identify potentially causal genes responsible for economic traits in Simmental beef cattle.J Anim Sci Biotechnol. 2023 May 11;14(1):78. doi: 10.1186/s40104-023-00876-7. J Anim Sci Biotechnol. 2023. PMID: 37165455 Free PMC article.
-
Association analyses of rare variants identify two genes associated with refractive error.PLoS One. 2022 Sep 22;17(9):e0272379. doi: 10.1371/journal.pone.0272379. eCollection 2022. PLoS One. 2022. PMID: 36137074 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

