Temperature-sensitive spinal muscular atrophy-causing point mutations lead to SMN instability, locomotor defects and premature lethality in Drosophila
- PMID: 32501283
- PMCID: PMC7325441
- DOI: 10.1242/dmm.043307
Temperature-sensitive spinal muscular atrophy-causing point mutations lead to SMN instability, locomotor defects and premature lethality in Drosophila
Abstract
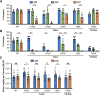
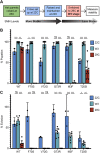
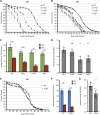
Spinal muscular atrophy (SMA) is the leading genetic cause of death in young children, arising from homozygous deletion or mutation of the survival motor neuron 1 (SMN1) gene. SMN protein expressed from a paralogous gene, SMN2, is the primary genetic modifier of SMA; small changes in overall SMN levels cause dramatic changes in disease severity. Thus, deeper insight into mechanisms that regulate SMN protein stability should lead to better therapeutic outcomes. Here, we show that SMA patient-derived missense mutations in the Drosophila SMN Tudor domain exhibit a pronounced temperature sensitivity that affects organismal viability, larval locomotor function and adult longevity. These disease-related phenotypes are domain specific and result from decreased SMN stability at elevated temperature. This system was utilized to manipulate SMN levels during various stages of Drosophila development. Owing to a large maternal contribution of mRNA and protein, Smn is not expressed zygotically during embryogenesis. Interestingly, we find that only baseline levels of SMN are required during larval stages, whereas high levels of the protein are required during pupation. This previously uncharacterized period of elevated SMN expression, during which the majority of adult tissues are formed and differentiated, could be an important and translationally relevant developmental stage in which to study SMN function. Taken together, these findings illustrate a novel in vivo role for the SMN Tudor domain in maintaining SMN homeostasis and highlight the necessity for high SMN levels at crucial developmental time points that are conserved from Drosophila to humans.
Keywords: Drosophila models of human disease; SMN protein; Spinal muscular atrophy; Tudor domain.
© 2020. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures




References
-
- Alatorre-Jiménez M., Sánchez-Luna J., González-Renovato E., Sánchez-López A., Sánchez-Luna S., Hernández-Navarro V., Pacheco-Moises F. and Ortiz G. (2015). Spinal muscular atrophy: review of a child onset disease. Br. J. Med. Med. Res. 6, 647-660. 10.9734/BJMMR/2015/14251 - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

