Temporal single-cell regeneration studies: the greatest thing since sliced pancreas?
- PMID: 34006411
- PMCID: PMC8239162
- DOI: 10.1016/j.tem.2021.04.009
Temporal single-cell regeneration studies: the greatest thing since sliced pancreas?
Abstract
The application of single-cell analytic techniques to the study of stem/progenitor cell niches supports the emerging view that pancreatic cell lineages are in a state of flux between differentiation stages. For all their value, however, such analyses merely offer a snapshot of the cellular palette of the tissue at any given time point. Conclusions about potential developmental/regeneration paths are solely based on bioinformatics inferences. In this context, the advent of new techniques for the long-term culture and lineage tracing of human pancreatic slices offers a virtual window into the native organ and presents the field with a unique opportunity to serially resolve pancreatic regeneration dynamics at the single-cell level.
Keywords: human pancreatic slices; pancreatic regeneration; pseudotime; single-cell RNA sequencing; β-cell neogenesis.
Copyright © 2021 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of interests J. D-B. is co-inventor of intellectual property owned by the University of Miami and described in this article, but receives no royalties in connection with it.
Figures
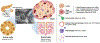

Similar articles
-
Dynamic scRNA-seq of live human pancreatic slices reveals functional endocrine cell neogenesis through an intermediate ducto-acinar stage.Cell Metab. 2023 Nov 7;35(11):1944-1960.e7. doi: 10.1016/j.cmet.2023.10.001. Epub 2023 Oct 27. Cell Metab. 2023. PMID: 37898119
-
Lineage tracing of pancreatic stem cells and beta cell regeneration.Methods Mol Biol. 2012;933:303-15. doi: 10.1007/978-1-62703-068-7_20. Methods Mol Biol. 2012. PMID: 22893416
-
Fate mapping of ptf1a-expressing cells during pancreatic organogenesis and regeneration in zebrafish.Dev Dyn. 2015 Jun;244(6):724-35. doi: 10.1002/dvdy.24271. Epub 2015 May 22. Dev Dyn. 2015. PMID: 25773748 Free PMC article.
-
Can pancreatic duct-derived progenitors be a source of islet regeneration?Biochem Biophys Res Commun. 2009 Jun 12;383(4):383-5. doi: 10.1016/j.bbrc.2009.03.114. Epub 2009 Mar 24. Biochem Biophys Res Commun. 2009. PMID: 19324022 Review.
-
β-Cell regeneration through the transdifferentiation of pancreatic cells: Pancreatic progenitor cells in the pancreas.J Diabetes Investig. 2016 May;7(3):286-96. doi: 10.1111/jdi.12475. Epub 2016 Feb 29. J Diabetes Investig. 2016. PMID: 27330712 Free PMC article. Review.
Cited by
-
Engineered tools to study endocrine dysfunction of pancreas.Biophys Rev (Melville). 2024 Oct 22;5(4):041303. doi: 10.1063/5.0220396. eCollection 2024 Dec. Biophys Rev (Melville). 2024. PMID: 39449867 Review.
-
Pancreatic β-cell regeneration in situ by the ALK3 agonist THR-123.Nat Commun. 2025 Jul 3;16(1):6121. doi: 10.1038/s41467-025-61534-2. Nat Commun. 2025. PMID: 40610457 Free PMC article.
-
Sex-specific regulatory architecture of pancreatic islets from subjects with and without type 2 diabetes.EMBO J. 2024 Dec;43(24):6364-6382. doi: 10.1038/s44318-024-00313-z. Epub 2024 Nov 20. EMBO J. 2024. PMID: 39567827 Free PMC article.
-
Single cell regulatory architecture of human pancreatic islets suggests sex differences in β cell function and the pathogenesis of type 2 diabetes.Res Sq [Preprint]. 2024 Jul 3:rs.3.rs-4607352. doi: 10.21203/rs.3.rs-4607352/v1. Res Sq. 2024. Update in: EMBO J. 2024 Dec;43(24):6364-6382. doi: 10.1038/s44318-024-00313-z. PMID: 39011095 Free PMC article. Updated. Preprint.
-
Single cell regulatory architecture of human pancreatic islets suggests sex differences in β cell function and the pathogenesis of type 2 diabetes.bioRxiv [Preprint]. 2024 Jun 11:2024.04.11.589096. doi: 10.1101/2024.04.11.589096. bioRxiv. 2024. Update in: EMBO J. 2024 Dec;43(24):6364-6382. doi: 10.1038/s44318-024-00313-z. PMID: 38645001 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

