Nuclear organisation and replication timing are coupled through RIF1-PP1 interaction
- PMID: 34006872
- PMCID: PMC8131703
- DOI: 10.1038/s41467-021-22899-2
Nuclear organisation and replication timing are coupled through RIF1-PP1 interaction
Abstract
Three-dimensional genome organisation and replication timing are known to be correlated, however, it remains unknown whether nuclear architecture overall plays an instructive role in the replication-timing programme and, if so, how. Here we demonstrate that RIF1 is a molecular hub that co-regulates both processes. Both nuclear organisation and replication timing depend upon the interaction between RIF1 and PP1. However, whereas nuclear architecture requires the full complement of RIF1 and its interaction with PP1, replication timing is not sensitive to RIF1 dosage. The role of RIF1 in replication timing also extends beyond its interaction with PP1. Availing of this separation-of-function approach, we have therefore identified in RIF1 dual function the molecular bases of the co-dependency of the replication-timing programme and nuclear architecture.
Conflict of interest statement
The authors declare no competing interests.
Figures
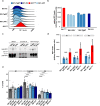
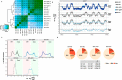
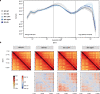

Similar articles
-
Nuclear Architecture Organized by Rif1 Underpins the Replication-Timing Program.Mol Cell. 2016 Jan 21;61(2):260-73. doi: 10.1016/j.molcel.2015.12.001. Epub 2015 Dec 24. Mol Cell. 2016. PMID: 26725008 Free PMC article.
-
Rif1 acts through Protein Phosphatase 1 but independent of replication timing to suppress telomere extension in budding yeast.Nucleic Acids Res. 2018 May 4;46(8):3993-4003. doi: 10.1093/nar/gky132. Nucleic Acids Res. 2018. PMID: 29529242 Free PMC article.
-
Mouse Rif1 is a regulatory subunit of protein phosphatase 1 (PP1).Sci Rep. 2017 May 18;7(1):2119. doi: 10.1038/s41598-017-01910-1. Sci Rep. 2017. PMID: 28522851 Free PMC article.
-
G-quadruplex binding protein Rif1, a key regulator of replication timing.J Biochem. 2021 Feb 6;169(1):1-14. doi: 10.1093/jb/mvaa128. J Biochem. 2021. PMID: 33169133 Review.
-
Rif1-Dependent Control of Replication Timing.Genes (Basel). 2022 Mar 20;13(3):550. doi: 10.3390/genes13030550. Genes (Basel). 2022. PMID: 35328102 Free PMC article. Review.
Cited by
-
DDK: The Outsourced Kinase of Chromosome Maintenance.Biology (Basel). 2022 Jun 7;11(6):877. doi: 10.3390/biology11060877. Biology (Basel). 2022. PMID: 35741398 Free PMC article. Review.
-
RIF1 integrates DNA repair and transcriptional requirements during the establishment of humoral immune responses.Nat Commun. 2025 Jan 17;16(1):777. doi: 10.1038/s41467-025-56166-5. Nat Commun. 2025. PMID: 39824820 Free PMC article.
-
Preventing excess replication origin activation to ensure genome stability.Trends Genet. 2022 Feb;38(2):169-181. doi: 10.1016/j.tig.2021.09.008. Epub 2021 Oct 6. Trends Genet. 2022. PMID: 34625299 Free PMC article. Review.
-
Dynamics of Replication-Associated Protein Levels through the Cell Cycle.Int J Mol Sci. 2024 Jul 28;25(15):8230. doi: 10.3390/ijms25158230. Int J Mol Sci. 2024. PMID: 39125800 Free PMC article.
-
A global high-density chromatin interaction network reveals functional long-range and trans-chromosomal relationships.Genome Biol. 2022 Nov 9;23(1):238. doi: 10.1186/s13059-022-02790-z. Genome Biol. 2022. PMID: 36352464 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

