Metaplasmidome-encoded functions of Siberian low-centered polygonal tundra soils
- PMID: 34012103
- PMCID: PMC8528913
- DOI: 10.1038/s41396-021-01003-y
Metaplasmidome-encoded functions of Siberian low-centered polygonal tundra soils
Abstract
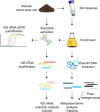
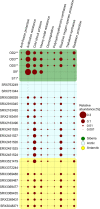
Plasmids have the potential to transfer genetic traits within bacterial communities and thereby serve as a crucial tool for the rapid adaptation of bacteria in response to changing environmental conditions. Our knowledge of the environmental pool of plasmids (the metaplasmidome) and encoded functions is still limited due to a lack of sufficient extraction methods and tools for identifying and assembling plasmids from metagenomic datasets. Here, we present the first insights into the functional potential of the metaplasmidome of permafrost-affected active-layer soil-an environment with a relatively low biomass and seasonal freeze-thaw cycles that is strongly affected by global warming. The obtained results were compared with plasmid-derived sequences extracted from polar metagenomes. Metaplasmidomes from the Siberian active layer were enriched via cultivation, which resulted in a longer contig length as compared with plasmids that had been directly retrieved from the metagenomes of polar environments. The predicted hosts of plasmids belonged to Moraxellaceae, Pseudomonadaceae, Enterobacteriaceae, Pectobacteriaceae, Burkholderiaceae, and Firmicutes. Analysis of their genetic content revealed the presence of stress-response genes, including antibiotic and metal resistance determinants, as well as genes encoding protectants against the cold.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Shifts of tundra bacterial and archaeal communities along a permafrost thaw gradient in Alaska.Mol Ecol. 2015 Jan;24(1):222-34. doi: 10.1111/mec.13015. Epub 2014 Dec 31. Mol Ecol. 2015. PMID: 25424441
-
Long-Term Warming in Alaska Enlarges the Diazotrophic Community in Deep Soils.mBio. 2019 Feb 26;10(1):e02521-18. doi: 10.1128/mBio.02521-18. mBio. 2019. PMID: 30808694 Free PMC article.
-
Contrasting above- and belowground organic matter decomposition and carbon and nitrogen dynamics in response to warming in High Arctic tundra.Glob Chang Biol. 2018 Jun;24(6):2660-2672. doi: 10.1111/gcb.14017. Epub 2018 Jan 7. Glob Chang Biol. 2018. PMID: 29235209
-
Dynamics of microbial communities and CO2 and CH4 fluxes in the tundra ecosystems of the changing Arctic.J Microbiol. 2019 May;57(5):325-336. doi: 10.1007/s12275-019-8661-2. Epub 2019 Jan 16. J Microbiol. 2019. PMID: 30656588 Review.
-
Plasmids foster diversification and adaptation of bacterial populations in soil.FEMS Microbiol Rev. 2012 Nov;36(6):1083-104. doi: 10.1111/j.1574-6976.2012.00337.x. Epub 2012 Mar 26. FEMS Microbiol Rev. 2012. PMID: 22393901 Review.
Cited by
-
Landscape of the metaplasmidome of deep-sea hydrothermal vents located at Arctic Mid-Ocean Ridges in the Norwegian-Greenland Sea: ecological insights from comparative analysis of plasmid identification tools.FEMS Microbiol Ecol. 2024 Sep 14;100(10):fiae124. doi: 10.1093/femsec/fiae124. FEMS Microbiol Ecol. 2024. PMID: 39271469 Free PMC article.
-
The Petri dish under the ice: permafrost pathogens and their impact on global healthcare and antibiotic resistance.Ann Med Surg (Lond). 2024 Oct 11;86(12):7193-7201. doi: 10.1097/MS9.0000000000002650. eCollection 2024 Dec. Ann Med Surg (Lond). 2024. PMID: 39649926 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

