This is a preprint.
Early cross-coronavirus reactive signatures of protective humoral immunity against COVID-19
- PMID: 34013263
- PMCID: PMC8132219
- DOI: 10.1101/2021.05.11.443609
Early cross-coronavirus reactive signatures of protective humoral immunity against COVID-19
Update in
-
Early cross-coronavirus reactive signatures of humoral immunity against COVID-19.Sci Immunol. 2021 Oct 15;6(64):eabj2901. doi: 10.1126/sciimmunol.abj2901. Epub 2021 Oct 15. Sci Immunol. 2021. PMID: 34652962 Free PMC article.
Abstract
The introduction of vaccines has inspired new hope in the battle against SARS-CoV-2. However, the emergence of viral variants, in the absence of potent antivirals, has left the world struggling with the uncertain nature of this disease. Antibodies currently represent the strongest correlate of immunity against COVID-19, thus we profiled the earliest humoral signatures in a large cohort of severe and asymptomatic COVID-19 individuals. While a SARS-CoV-2-specific immune response evolved rapidly in survivors of COVID-19, non-survivors exhibited blunted and delayed humoral immune evolution, particularly with respect to S2-specific antibody evolution. Given the conservation of S2 across β-coronaviruses, we found the early development of SARS-CoV-2-specific immunity occurred in tandem with pre-existing common β-coronavirus OC43 humoral immunity in survivors, which was selectively also expanded in individuals that develop paucisymptomatic infection. These data point to the importance of cross-coronavirus immunity as a correlate of protection against COVID-19.
Conflict of interest statement
Conflict of interests:
G.A. is a founder of Seromyx Systems Inc. D.D., P.M., A.S.M, and E.R.M. are employees of Space Exploration Technologies Corp. E.S.F. is a founder, scientific advisor and equity holder for: Jengu Therapeutics (board member), Neomorph Inc and Civetta Therapeutics; an equity holder in C4 Therapeutics (CCCC); and a consultant to Novartis, Sanofi, AbbVie, Pfizer, Astellas, EcoR1 capital and Deerfield. The Fischer lab receives or has received research funding from Novartis, Ajax, and Astellas not related to this work. All other authors have declared that no conflict of interest exists.
Figures



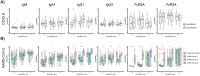
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
