Abortive intussusceptive angiogenesis causes multi-cavernous vascular malformations
- PMID: 34013885
- PMCID: PMC8175082
- DOI: 10.7554/eLife.62155
Abortive intussusceptive angiogenesis causes multi-cavernous vascular malformations
Abstract
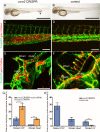
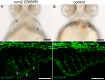
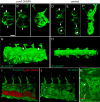
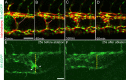
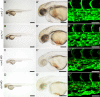
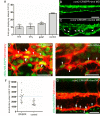
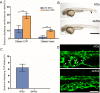
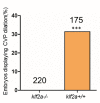
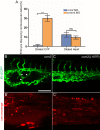
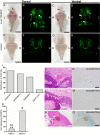
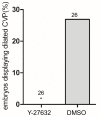
Mosaic inactivation of CCM2 in humans causes cerebral cavernous malformations (CCMs) containing adjacent dilated blood-filled multi-cavernous lesions. We used CRISPR-Cas9 mutagenesis to induce mosaic inactivation of zebrafish ccm2 resulting in a novel lethal multi-cavernous lesion in the embryonic caudal venous plexus (CVP) caused by obstruction of blood flow by intraluminal pillars. These pillars mimic those that mediate intussusceptive angiogenesis; however, in contrast to the normal process, the pillars failed to fuse to split the pre-existing vessel in two. Abortive intussusceptive angiogenesis stemmed from mosaic inactivation of ccm2 leading to patchy klf2a overexpression and resultant aberrant flow signaling. Surviving adult fish manifested histologically typical hemorrhagic CCM. Formation of mammalian CCM requires the flow-regulated transcription factor KLF2; fish CCM and the embryonic CVP lesion failed to form in klf2a null fish indicating a common pathogenesis with the mammalian lesion. These studies describe a zebrafish CCM model and establish a mechanism that can explain the formation of characteristic multi-cavernous lesions.
Keywords: angiogenesis; blood flow signaling; cerebral cavernous malformations; developmental biology; zebrafish.
© 2021, Li et al.
Conflict of interest statement
WL, VT, IS, BX, TM, RL, DK, IA, MG No competing interests declared
Figures





References
-
- Boulday G, Rudini N, Maddaluno L, Blécon A, Arnould M, Gaudric A, Chapon F, Adams RH, Dejana E, Tournier-Lasserve E. Developmental timing of CCM2 loss influences cerebral cavernous malformations in mice. Journal of Experimental Medicine. 2011;208:1835–1847. doi: 10.1084/jem.20110571. - DOI - PMC - PubMed
-
- Chan AC, Drakos SG, Ruiz OE, Smith ACH, Gibson CC, Ling J, Passi SF, Stratman AN, Sacharidou A, Revelo MP, Grossmann AH, Diakos NA, Davis GE, Metzstein MM, Whitehead KJ, Li DY. Mutations in 2 distinct genetic pathways result in cerebral cavernous malformations in mice. Journal of Clinical Investigation. 2011;121:1871–1881. doi: 10.1172/JCI44393. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

