Remarkable and Unexpected Mechanism for (S)-3-Amino-4-(difluoromethylenyl)cyclohex-1-ene-1-carboxylic Acid as a Selective Inactivator of Human Ornithine Aminotransferase
- PMID: 34014654
- PMCID: PMC8369387
- DOI: 10.1021/jacs.1c03572
Remarkable and Unexpected Mechanism for (S)-3-Amino-4-(difluoromethylenyl)cyclohex-1-ene-1-carboxylic Acid as a Selective Inactivator of Human Ornithine Aminotransferase
Abstract
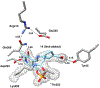
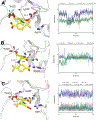
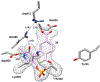
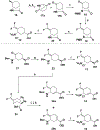
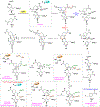
Human ornithine aminotransferase (hOAT) is a pyridoxal 5'-phosphate (PLP)-dependent enzyme that was recently found to play an important role in the metabolic reprogramming of hepatocellular carcinoma (HCC) via the proline and glutamine metabolic pathways. The selective inhibition of hOAT by compound 10 exhibited potent in vivo antitumor activity. Inspired by the discovery of the aminotransferase inactivator (1S,3S)-3-amino-4-(difluoromethylene)cyclopentane-1-carboxylic acid (5), we rationally designed, synthesized, and evaluated a series of six-membered-ring analogs. Among them, 14 was identified as a new selective hOAT inactivator, which demonstrated a potency 22× greater than that of 10. Three different types of protein mass spectrometry approaches and two crystallographic approaches were employed to identify the structure of hOAT-14 and the formation of a remarkable final adduct (32') in the active site. These spectral studies reveal an enzyme complex heretofore not observed in a PLP-dependent enzyme, which has covalent bonds to two nearby residues. Crystal soaking experiments and molecular dynamics simulations were carried out to identify the structure of the active-site intermediate 27' and elucidate the order of the two covalent bonds that formed, leading to 32'. The initial covalent reaction of the activated warhead occurs with *Thr322 from the second subunit, followed by a subsequent nucleophilic attack by the catalytic residue Lys292. The turnover mechanism of 14 by hOAT was supported by a mass spectrometric analysis of metabolites and fluoride ion release experiments. This novel mechanism for hOAT with 14 will contribute to the further rational design of selective inactivators and an understanding of potential inactivation mechanisms by aminotransferases.
Figures



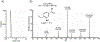
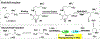



Similar articles
-
Design, Synthesis, and Mechanistic Studies of (R)-3-Amino-5,5-difluorocyclohex-1-ene-1-carboxylic Acid as an Inactivator of Human Ornithine Aminotransferase.ACS Chem Biol. 2024 May 17;19(5):1066-1081. doi: 10.1021/acschembio.4c00022. Epub 2024 Apr 17. ACS Chem Biol. 2024. PMID: 38630468 Free PMC article.
-
A Remarkable Difference That One Fluorine Atom Confers on the Mechanisms of Inactivation of Human Ornithine Aminotransferase by Two Cyclohexene Analogues of γ-Aminobutyric Acid.J Am Chem Soc. 2020 Mar 11;142(10):4892-4903. doi: 10.1021/jacs.0c00193. Epub 2020 Mar 1. J Am Chem Soc. 2020. PMID: 32114761 Free PMC article.
-
Rational Design, Synthesis, and Mechanism of (3S,4R)-3-Amino-4-(difluoromethyl)cyclopent-1-ene-1-carboxylic Acid: Employing a Second-Deprotonation Strategy for Selectivity of Human Ornithine Aminotransferase over GABA Aminotransferase.J Am Chem Soc. 2022 Mar 30;144(12):5629-5642. doi: 10.1021/jacs.2c00924. Epub 2022 Mar 16. J Am Chem Soc. 2022. PMID: 35293728 Free PMC article.
-
Ornithine aminotransferase versus GABA aminotransferase: implications for the design of new anticancer drugs.Med Res Rev. 2015 Mar;35(2):286-305. doi: 10.1002/med.21328. Epub 2014 Aug 22. Med Res Rev. 2015. PMID: 25145640 Free PMC article. Review.
-
Design of potential anticonvulsant agents: mechanistic classification of GABA aminotransferase inactivators.J Med Chem. 1989 Nov;32(11):2413-21. doi: 10.1021/jm00131a001. J Med Chem. 1989. PMID: 2681782 Review.
Cited by
-
Synthesis of (2R,4S)-4-Amino-5-hydroxybicyclo[3.1.1]heptane-2-carboxylic Acid via an Asymmetric Intramolecular Mannich Reaction.J Org Chem. 2024 Jun 21;89(12):9110-9117. doi: 10.1021/acs.joc.4c00781. Epub 2024 Jun 10. J Org Chem. 2024. PMID: 38857432 Free PMC article.
-
Inactivators of Ornithine Aminotransferase for the Treatment of Hepatocellular Carcinoma.ACS Med Chem Lett. 2021 Dec 9;13(1):38-49. doi: 10.1021/acsmedchemlett.1c00526. eCollection 2022 Jan 13. ACS Med Chem Lett. 2021. PMID: 35059122 Free PMC article.
-
Information-Rich, Dual-Function 13C/2H-Isotopic Crosstalk NMR Assay for Human Serine Racemase (hSR) Provides a PLP-Enzyme "Partitioning Fingerprint" and Reveals Disparate Chemotypes for hSR Inhibition.J Am Chem Soc. 2023 Feb 8;145(5):3158-3174. doi: 10.1021/jacs.2c12774. Epub 2023 Jan 25. J Am Chem Soc. 2023. PMID: 36696670 Free PMC article.
-
α-Hydrazino Acids Inhibit Pyridoxal Phosphate-Dependent Decarboxylases via "Catalytically Correct" Ketoenamine Tautomers: A Special Motif for Chemical Biology and Drug Discovery?ACS Catal. 2025 May 2;15(10):8204-8218. doi: 10.1021/acscatal.5c00326. eCollection 2025 May 16. ACS Catal. 2025. PMID: 40401103
-
Determination of the pH dependence, substrate specificity, and turnovers of alternative substrates for human ornithine aminotransferase.J Biol Chem. 2022 Jun;298(6):101969. doi: 10.1016/j.jbc.2022.101969. Epub 2022 Apr 20. J Biol Chem. 2022. PMID: 35460691 Free PMC article.
References
-
- Sayiner M; Golabi P; Younossi ZM Disease burden of hepatocellular carcinoma: a global perspective. Digest Dis. Sci 2019, 64, 910–917. - PubMed
-
- Personeni N; Rimassa L. Hepatocellular carcinoma: a global disease in need of individualized treatment strategies. J. Oncol. Pract 2017, 13, 368–370. - PubMed
-
- Sherman M; Bruix J; Porayko M; Tran T; Comm APG Screening for hepatocellular carcinoma: The rationale for the American association for the study of liver diseases recommendations. Hepatology 2012, 56, 793–796. - PubMed
-
- Leathers JS; Balderramo D; Prieto J; Diehl F; Gonzalez-Ballerga E; Ferreiro MR; Carrera E; Barreyro F; Diaz-Ferrer J; Singh D; Mattos AZ; Carrilho F; Debes JD Sorafenib for treatment of hepatocellular carcinoma a survival analysis from the aouth american liver research network. J. Clin. Gastroenterol 2019, 53, 464–469. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

