The expanding field of non-canonical RNA capping: new enzymes and mechanisms
- PMID: 34017598
- PMCID: PMC8131947
- DOI: 10.1098/rsos.201979
The expanding field of non-canonical RNA capping: new enzymes and mechanisms
Abstract
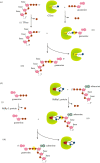
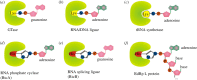
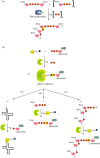
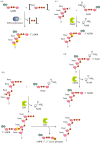
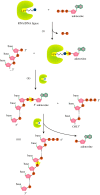
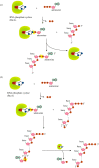
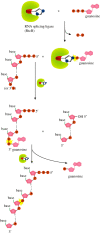
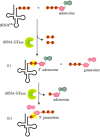
Recent years witnessed the discovery of ubiquitous and diverse 5'-end RNA cap-like modifications in prokaryotes as well as in eukaryotes. These non-canonical caps include metabolic cofactors, such as NAD+/NADH, FAD, cell wall precursors UDP-GlcNAc, alarmones, e.g. dinucleotides polyphosphates, ADP-ribose and potentially other nucleoside derivatives. They are installed at the 5' position of RNA via template-dependent incorporation of nucleotide analogues as an initiation substrate by RNA polymerases. However, the discovery of NAD-capped processed RNAs in human cells suggests the existence of alternative post-transcriptional NC capping pathways. In this review, we compiled growing evidence for a number of these other mechanisms which produce various non-canonically capped RNAs and a growing repertoire of capping small molecules. Enzymes shown to be involved are ADP-ribose polymerases, glycohydrolases and tRNA synthetases, and may potentially include RNA 3'-phosphate cyclases, tRNA guanylyl transferases, RNA ligases and ribozymes. An emerging rich variety of capping molecules and enzymes suggests an unrecognized level of complexity of RNA metabolism.
Keywords: RNA processing; capping; non-canonical capping.
© 2021 The Authors.
Figures

References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
