Adaptive Proteome Diversification by Nonsynonymous A-to-I RNA Editing in Coleoid Cephalopods
- PMID: 34022057
- PMCID: PMC8382921
- DOI: 10.1093/molbev/msab154
Adaptive Proteome Diversification by Nonsynonymous A-to-I RNA Editing in Coleoid Cephalopods
Abstract
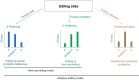
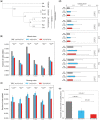
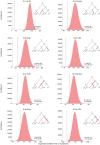
RNA editing by the ADAR enzymes converts selected adenosines into inosines, biological mimics for guanosines. By doing so, it alters protein-coding sequences, resulting in novel protein products that diversify the proteome beyond its genomic blueprint. Recoding is exceptionally abundant in the neural tissues of coleoid cephalopods (octopuses, squids, and cuttlefishes), with an over-representation of nonsynonymous edits suggesting positive selection. However, the extent to which proteome diversification by recoding provides an adaptive advantage is not known. It was recently suggested that the role of evolutionarily conserved edits is to compensate for harmful genomic substitutions, and that there is no added value in having an editable codon as compared with a restoration of the preferred genomic allele. Here, we show that this hypothesis fails to explain the evolutionary dynamics of recoding sites in coleoids. Instead, our results indicate that a large fraction of the shared, strongly recoded, sites in coleoids have been selected for proteome diversification, meaning that the fitness of an editable A is higher than an uneditable A or a genomically encoded G.
Keywords: RNA editing; adaptation; evolution.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures

Similar articles
-
The preponderance of nonsynonymous A-to-I RNA editing in coleoids is nonadaptive.Nat Commun. 2019 Nov 27;10(1):5411. doi: 10.1038/s41467-019-13275-2. Nat Commun. 2019. PMID: 31776345 Free PMC article.
-
High-level RNA editing diversifies the coleoid cephalopod brain proteome.Brief Funct Genomics. 2023 Nov 17;22(6):525-532. doi: 10.1093/bfgp/elad034. Brief Funct Genomics. 2023. PMID: 37981860
-
Extensive Recoding of the Neural Proteome in Cephalopods by RNA Editing.Annu Rev Anim Biosci. 2023 Feb 15;11:57-75. doi: 10.1146/annurev-animal-060322-114534. Annu Rev Anim Biosci. 2023. PMID: 36790891 Review.
-
Trade-off between Transcriptome Plasticity and Genome Evolution in Cephalopods.Cell. 2017 Apr 6;169(2):191-202.e11. doi: 10.1016/j.cell.2017.03.025. Cell. 2017. PMID: 28388405 Free PMC article.
-
Proteome Diversification by RNA Editing.Methods Mol Biol. 2021;2181:229-251. doi: 10.1007/978-1-0716-0787-9_14. Methods Mol Biol. 2021. PMID: 32729084 Review.
Cited by
-
Adaptive evolution of A-to-I auto-editing site in Adar of eusocial insects.BMC Genomics. 2024 Aug 26;25(1):803. doi: 10.1186/s12864-024-10709-0. BMC Genomics. 2024. PMID: 39187830 Free PMC article.
-
Autorecoding A-to-I RNA editing sites in the Adar gene underwent compensatory gains and losses in major insect clades.RNA. 2023 Oct;29(10):1509-1519. doi: 10.1261/rna.079682.123. Epub 2023 Jul 14. RNA. 2023. PMID: 37451866 Free PMC article.
-
Adaptation of A-to-I RNA editing in bacteria, fungi, and animals.Front Microbiol. 2023 May 24;14:1204080. doi: 10.3389/fmicb.2023.1204080. eCollection 2023. Front Microbiol. 2023. PMID: 37293227 Free PMC article. No abstract available.
-
MicroRNAs are deeply linked to the emergence of the complex octopus brain.Sci Adv. 2022 Nov 25;8(47):eadd9938. doi: 10.1126/sciadv.add9938. Epub 2022 Nov 25. Sci Adv. 2022. PMID: 36427315 Free PMC article.
-
A hierarchy in clusters of cephalopod mRNA editing sites.Sci Rep. 2022 Mar 2;12(1):3447. doi: 10.1038/s41598-022-07460-5. Sci Rep. 2022. PMID: 35236910 Free PMC article.
References
-
- Anderson FE, Lindgren AR.. 2021. Phylogenomic analyses recover a clade of large-bodied decapodiform cephalopods. Mol Phylogenet Evol. 156:107038. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

