An endangered flightless grasshopper with strong genetic structure maintains population genetic variation despite extensive habitat loss
- PMID: 34026013
- PMCID: PMC8131777
- DOI: 10.1002/ece3.7428
An endangered flightless grasshopper with strong genetic structure maintains population genetic variation despite extensive habitat loss
Abstract
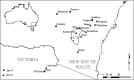
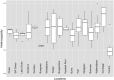
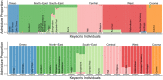
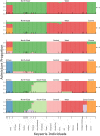
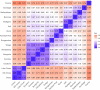
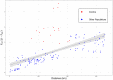
Conservation research is dominated by vertebrate examples but the shorter generation times and high local population sizes of invertebrates may lead to very different management strategies, particularly for species with low movement rates. Here we investigate the genetic structure of an endangered flightless grasshopper, Keyacris scurra, which was used in classical evolutionary studies in the 1960s. It had a wide distribution across New South Wales (NSW) and Victoria in pre-European times but has now become threatened because of land clearing for agriculture and other activities. We revisited remnant sites of K. scurra, with populations now restricted to only one area in Victoria and a few small patches in NSW and the Australian Capital Territory (ACT). Using DArtseq to generate SNP markers as well as mtDNA sequence data, we show that the remaining Victorian populations in an isolated valley are genetically distinct from the NSW populations and that all populations tend to be genetically unique, with large F ST values up to 0.8 being detected for the SNP datasets. We also find that, with one notable exception, the NSW/ACT populations separate genetically into previously described chromosomal races (2n = 15 vs. 2n = 17). Isolation by distance was detected across both the SNP and mtDNA datasets, and there was substantial differentiation within chromosomal races. Genetic diversity as measured by heterozygosity was not correlated with the size of remaining habitat where the populations were found, with high variation present in some remnant cemetery sites. However, inbreeding correlated negatively with estimated habitat size at 25-500 m patch radius. These findings emphasize the importance of small habitat areas in conserving genetic variation in such species with low mobility, and they highlight populations suitable for future translocation efforts.
Keywords: Keyacris; fragmentation; grassland; isolation by distance; morabine; small population area.
© 2021 The Authors. Ecology and Evolution published by John Wiley & Sons Ltd.
Conflict of interest statement
None declared.
Figures

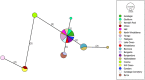



References
-
- Allard, R. , & Wehrhahn, C. (1964). A theory which predicts stable equilibrium for inversion polymorphisms in the grasshopper Moraba scurra . Evolution, 18, 129–130.
-
- Allendorf, F. W. , Hohenlohe, P. A. , & Luikart, G. (2010). Genomics and the future of conservation genetics. Nature Reviews Genetics, 11, 697–709. - PubMed
-
- Black, S. H. , & Vaughan, M. (2009). Endangered Insects. Encyclopedia of Insects (Second Edition). Elsevier.
-
- Blackith, R. , & Blackith, R. M. (1969). Observations on the biology of some morabine grasshoppers. Australian Journal of Zoology, 17, 1–12.
LinkOut - more resources
Full Text Sources
Other Literature Sources

