Single-Entity Detection With TEM-Fabricated Nanopores
- PMID: 34026729
- PMCID: PMC8138203
- DOI: 10.3389/fchem.2021.664820
Single-Entity Detection With TEM-Fabricated Nanopores
Abstract
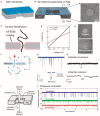
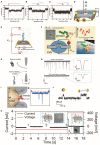
Nanopore-based single-entity detection shows immense potential in sensing and sequencing technologies. Solid-state nanopores permit unprecedented detail while preserving mechanical robustness, reusability, adjustable pore size, and stability in different physical and chemical environments. The transmission electron microscope (TEM) has evolved into a powerful tool for fabricating and characterizing nanometer-sized pores within a solid-state ultrathin membrane. By detecting differences in the ionic current signals due to single-entity translocation through the nanopore, solid-state nanopores can enable gene sequencing and single molecule/nanoparticle detection with high sensitivity, improved acquisition speed, and low cost. Here we briefly discuss the recent progress in the modification and characterization of TEM-fabricated nanopores. Moreover, we highlight some key applications of these nanopores in nucleic acids, protein, and nanoparticle detection. Additionally, we discuss the future of computer simulations in DNA and protein sequencing strategies. We also attempt to identify the challenges and discuss the future development of nanopore-detection technology aiming to promote the next-generation sequencing technology.
Keywords: TEM fabrication; electron-beam drilling; sequencing; single entity detection; solid-state nanopores.
Copyright © 2021 Yang, Saqib and Hao.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
TEM based applications in solid state nanopores: From fabrication to liquid in-situ bio-imaging.Micron. 2022 Nov;162:103347. doi: 10.1016/j.micron.2022.103347. Epub 2022 Sep 1. Micron. 2022. PMID: 36081256 Review.
-
Challenges of Single-Molecule DNA Sequencing with Solid-State Nanopores.Adv Exp Med Biol. 2019;1129:131-142. doi: 10.1007/978-981-13-6037-4_9. Adv Exp Med Biol. 2019. PMID: 30968365 Review.
-
Simple Fabrication of Solid-State Nanopores on a Carbon Film.Micromachines (Basel). 2021 Sep 21;12(9):1135. doi: 10.3390/mi12091135. Micromachines (Basel). 2021. PMID: 34577778 Free PMC article.
-
Fabrication and Applications of Solid-State Nanopores.Sensors (Basel). 2019 Apr 20;19(8):1886. doi: 10.3390/s19081886. Sensors (Basel). 2019. PMID: 31010038 Free PMC article. Review.
-
Electrochemical Reaction in Single Layer MoS2: Nanopores Opened Atom by Atom.Nano Lett. 2015 May 13;15(5):3431-8. doi: 10.1021/acs.nanolett.5b00768. Epub 2015 May 4. Nano Lett. 2015. PMID: 25928894
Cited by
-
Nanoscale Luminescence Imaging/Detection of Single Particles: State-of-the-Art and Future Prospects.ACS Meas Sci Au. 2023 Dec 7;4(1):3-24. doi: 10.1021/acsmeasuresciau.3c00052. eCollection 2024 Feb 21. ACS Meas Sci Au. 2023. PMID: 38404493 Free PMC article. Review.
-
Large-scale production of polyimide micropore-based flow cells for detecting nano-sized particles in fluids.RSC Adv. 2023 Jan 4;13(2):873-880. doi: 10.1039/d2ra07423k. eCollection 2023 Jan 3. RSC Adv. 2023. PMID: 36686911 Free PMC article.
-
Wafer-scale fabrication of solid-state nanopore array with a novel SpacerX process.Microsyst Nanoeng. 2025 Jun 25;11(1):129. doi: 10.1038/s41378-025-00979-3. Microsyst Nanoeng. 2025. PMID: 40555711 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

