The Rab7 effector WDR91 promotes autophagy-lysosome degradation in neurons by regulating lysosome fusion
- PMID: 34028500
- PMCID: PMC8150682
- DOI: 10.1083/jcb.202007061
The Rab7 effector WDR91 promotes autophagy-lysosome degradation in neurons by regulating lysosome fusion
Abstract
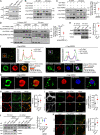
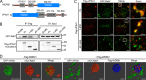
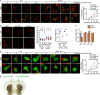
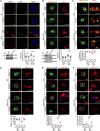
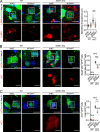
The effectors of the Rab7 small GTPase play multiple roles in Rab7-dependent endosome-lysosome and autophagy-lysosome pathways. However, it is largely unknown how distinct Rab7 effectors coordinate to maintain the homeostasis of late endosomes and lysosomes to ensure appropriate endolysosomal and autolysosomal degradation. Here we report that WDR91, a Rab7 effector required for early-to-late endosome conversion, is essential for lysosome function and homeostasis. Mice lacking Wdr91 specifically in the central nervous system exhibited behavioral defects and marked neuronal loss in the cerebral and cerebellar cortices. At the cellular level, WDR91 deficiency causes PtdIns3P-independent enlargement and dysfunction of lysosomes, leading to accumulation of autophagic cargoes in mouse neurons. WDR91 competes with the VPS41 subunit of the HOPS complex, another Rab7 effector, for binding to Rab7, thereby facilitating Rab7-dependent lysosome fusion in a controlled manner. WDR91 thus maintains an appropriate level of lysosome fusion to guard the normal function and survival of neurons.
© 2021 Xing et al.
Figures


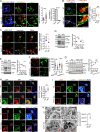
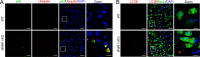

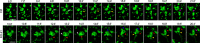
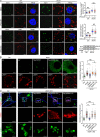
Similar articles
-
WDR91 is a Rab7 effector required for neuronal development.J Cell Biol. 2017 Oct 2;216(10):3307-3321. doi: 10.1083/jcb.201705151. Epub 2017 Aug 31. J Cell Biol. 2017. PMID: 28860274 Free PMC article.
-
Dynein Is Required for Rab7-Dependent Endosome Maturation, Retrograde Dendritic Transport, and Degradation.J Neurosci. 2022 Jun 1;42(22):4415-4434. doi: 10.1523/JNEUROSCI.2530-21.2022. Epub 2022 Apr 26. J Neurosci. 2022. PMID: 35474277 Free PMC article.
-
Phosphatidylinositol 4,5-bisphosphate controls Rab7 and PLEKHM1 membrane cycling during autophagosome-lysosome fusion.EMBO J. 2019 Apr 15;38(8):e100312. doi: 10.15252/embj.2018100312. Epub 2019 Mar 13. EMBO J. 2019. PMID: 31368593 Free PMC article.
-
Rab GTPase Function in Endosome and Lysosome Biogenesis.Trends Cell Biol. 2018 Nov;28(11):957-970. doi: 10.1016/j.tcb.2018.06.007. Epub 2018 Jul 17. Trends Cell Biol. 2018. PMID: 30025982 Review.
-
CORVET and HOPS tethering complexes - coordinators of endosome and lysosome fusion.J Cell Sci. 2013 Mar 15;126(Pt 6):1307-16. doi: 10.1242/jcs.107805. J Cell Sci. 2013. PMID: 23645161 Review.
Cited by
-
TRAF6 triggers Mycobacterium-infected host autophagy through Rab7 ubiquitination.Cell Death Discov. 2023 Nov 28;9(1):427. doi: 10.1038/s41420-023-01731-4. Cell Death Discov. 2023. PMID: 38016969 Free PMC article.
-
Exploring lysosomal biology: current approaches and methods.Biophys Rep. 2024 Apr 30;10(2):111-120. doi: 10.52601/bpr.2023.230028. Biophys Rep. 2024. PMID: 38774350 Free PMC article.
-
RAB22A sorts epithelial growth factor receptor (EGFR) from early endosomes to recycling endosomes for microvesicles release.J Extracell Vesicles. 2024 Jul;13(7):e12494. doi: 10.1002/jev2.12494. J Extracell Vesicles. 2024. PMID: 39051763 Free PMC article.
-
The role of lipophagy in liver cancer: mechanisms and targeted therapeutic interventions.Front Cell Dev Biol. 2025 Jul 8;13:1562542. doi: 10.3389/fcell.2025.1562542. eCollection 2025. Front Cell Dev Biol. 2025. PMID: 40698038 Free PMC article. Review.
-
Atractylenolide I inhibits angiogenesis and reverses sunitinib resistance in clear cell renal cell carcinoma through ATP6V0D2-mediated autophagic degradation of EPAS1/HIF2α.Autophagy. 2025 Mar;21(3):619-638. doi: 10.1080/15548627.2024.2421699. Epub 2024 Nov 4. Autophagy. 2025. PMID: 39477683 Free PMC article.
References
-
- Bröcker, C., Kuhlee A., Gatsogiannis C., Balderhaar H.J.K., Hönscher C., Engelbrecht-Vandré S., Ungermann C., and Raunser S.. 2012. Molecular architecture of the multisubunit homotypic fusion and vacuole protein sorting (HOPS) tethering complex. Proc. Natl. Acad. Sci. USA. 109:1991–1996. 10.1073/pnas.1117797109 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

