Wearable sensors enable personalized predictions of clinical laboratory measurements
- PMID: 34031607
- PMCID: PMC8293303
- DOI: 10.1038/s41591-021-01339-0
Wearable sensors enable personalized predictions of clinical laboratory measurements
Abstract
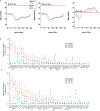
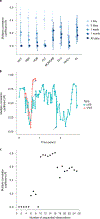
Vital signs, including heart rate and body temperature, are useful in detecting or monitoring medical conditions, but are typically measured in the clinic and require follow-up laboratory testing for more definitive diagnoses. Here we examined whether vital signs as measured by consumer wearable devices (that is, continuously monitored heart rate, body temperature, electrodermal activity and movement) can predict clinical laboratory test results using machine learning models, including random forest and Lasso models. Our results demonstrate that vital sign data collected from wearables give a more consistent and precise depiction of resting heart rate than do measurements taken in the clinic. Vital sign data collected from wearables can also predict several clinical laboratory measurements with lower prediction error than predictions made using clinically obtained vital sign measurements. The length of time over which vital signs are monitored and the proximity of the monitoring period to the date of prediction play a critical role in the performance of the machine learning models. These results demonstrate the value of commercial wearable devices for continuous and longitudinal assessment of physiological measurements that today can be measured only with clinical laboratory tests.
Conflict of interest statement
Competing interests
M.P.S. is a cofounder of Personalis, SensOmics, Qbio, January AI, Filtricine, Protos, and NiMo, and is on the scientific advisory board of Personalis, SensOmics, Qbio, January AI, Filtricine, Protos, NiMo, and Genapsys. All other authors have no competing interests.
Figures
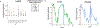





Comment in
-
Smartwatch biomarkers and the path to clinical use.Med. 2021 Jul 9;2(7):797-799. doi: 10.1016/j.medj.2021.06.005. Med. 2021. PMID: 35590218
References
-
- Sackett DL The rational clinical examination. A primer on the precision and accuracy of the clinical examination. J. Am. Med. Assoc 267, 2638–2644 (1992). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

