Activation of Cytochrome C Peroxidase Function Through Coordinated Foldon Loop Dynamics upon Interaction with Anionic Lipids
- PMID: 34033821
- PMCID: PMC8380053
- DOI: 10.1016/j.jmb.2021.167057
Activation of Cytochrome C Peroxidase Function Through Coordinated Foldon Loop Dynamics upon Interaction with Anionic Lipids
Abstract
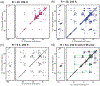
Cardiolipin (CL) is a mitochondrial anionic lipid that plays important roles in the regulation and signaling of mitochondrial apoptosis. CL peroxidation catalyzed by the assembly of CL-cytochrome c (cyt c) complexes at the inner mitochondrial membrane is a critical checkpoint. The structural changes in the protein, associated with peroxidase activation by CL and different anionic lipids, are not known at a molecular level. To better understand these peripheral protein-lipid interactions, we compare how phosphatidylglycerol (PG) and CL lipids trigger cyt c peroxidase activation, and correlate functional differences to structural and motional changes in membrane-associated cyt c. Structural and motional studies of the bound protein are enabled by magic angle spinning solid state NMR spectroscopy, while lipid peroxidase activity is assayed by mass spectrometry. PG binding results in a surface-bound state that preserves a nativelike fold, which nonetheless allows for significant peroxidase activity, though at a lower level than binding its native substrate CL. Lipid-specific differences in peroxidase activation are found to correlate to corresponding differences in lipid-induced protein mobility, affecting specific protein segments. The dynamics of omega loops C and D are upregulated by CL binding, in a way that is remarkably controlled by the protein:lipid stoichiometry. In contrast to complete chemical denaturation, membrane-induced protein destabilization reflects a destabilization of select cyt c foldons, while the energetically most stable helices are preserved. Our studies illuminate the interplay of protein and lipid dynamics in the creation of lipid peroxidase-active proteolipid complexes implicated in early stages of mitochondrial apoptosis.
Keywords: lipid peroxidation; lipidomics; mitochondrial apoptosis; peripheral membrane proteins; solid-state NMR spectroscopy.
Copyright © 2021 The Author(s). Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

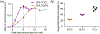



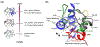
Similar articles
-
Structural Changes and Proapoptotic Peroxidase Activity of Cardiolipin-Bound Mitochondrial Cytochrome c.Biophys J. 2015 Nov 3;109(9):1873-84. doi: 10.1016/j.bpj.2015.09.016. Biophys J. 2015. PMID: 26536264 Free PMC article.
-
Surface-Binding to Cardiolipin Nanodomains Triggers Cytochrome c Pro-apoptotic Peroxidase Activity via Localized Dynamics.Structure. 2019 May 7;27(5):806-815.e4. doi: 10.1016/j.str.2019.02.007. Epub 2019 Mar 14. Structure. 2019. PMID: 30879887 Free PMC article.
-
Impact of Cardiolipin and Phosphatidylcholine Interactions on the Conformational Ensemble of Cytochrome c.Biochemistry. 2019 Aug 27;58(34):3617-3626. doi: 10.1021/acs.biochem.9b00495. Epub 2019 Aug 12. Biochemistry. 2019. PMID: 31380624
-
The "pro-apoptotic genies" get out of mitochondria: oxidative lipidomics and redox activity of cytochrome c/cardiolipin complexes.Chem Biol Interact. 2006 Oct 27;163(1-2):15-28. doi: 10.1016/j.cbi.2006.04.019. Epub 2006 May 12. Chem Biol Interact. 2006. PMID: 16797512 Review.
-
Structural transformations of cytochrome c upon interaction with cardiolipin.Chem Phys Lipids. 2014 Apr;179:57-63. doi: 10.1016/j.chemphyslip.2013.11.002. Epub 2013 Nov 16. Chem Phys Lipids. 2014. PMID: 24252639 Free PMC article. Review.
Cited by
-
Effect of COVID-19 mRNA Vaccine on Human Lung Carcinoma Cells In Vitro by Means of Raman Spectroscopy and Imaging.ACS Omega. 2023 Oct 30;8(45):42555-42564. doi: 10.1021/acsomega.3c05287. eCollection 2023 Nov 14. ACS Omega. 2023. PMID: 38024689 Free PMC article.
-
Anomalous peroxidase activity of cytochrome c is the primary pathogenic target in Barth syndrome.Nat Metab. 2023 Dec;5(12):2184-2205. doi: 10.1038/s42255-023-00926-4. Epub 2023 Nov 23. Nat Metab. 2023. PMID: 37996701 Free PMC article.
-
Order-to-Disorder and Disorder-to-Order Transitions of Proteins upon Binding to Phospholipid Membranes: Common Ground and Dissimilarities.Biomolecules. 2025 Jan 30;15(2):198. doi: 10.3390/biom15020198. Biomolecules. 2025. PMID: 40001501 Free PMC article. Review.
-
The plastid-encoded protein Orf2971 is required for protein translocation and chloroplast quality control.Plant Cell. 2022 Aug 25;34(9):3383-3399. doi: 10.1093/plcell/koac180. Plant Cell. 2022. PMID: 35708659 Free PMC article.
-
Solid-state NMR protocols for unveiling dynamics and (drug) interactions of membrane-bound proteins.Protein Sci. 2025 Apr;34(4):e70102. doi: 10.1002/pro.70102. Protein Sci. 2025. PMID: 40099898 Free PMC article.
References
-
- Alvarez-Paggi D, Hannibal L, Castro MA, Oviedo-Rouco S, Demicheli V, Tortora V, et al.Multifunctional cytochrome c: Learning new tricks from an old dog. Chemical Reviews. 2017;117:13382–460. - PubMed
-
- Ow YP, Green DR, Hao Z, Mak TW. Cytochrome c: functions beyond respiration. Nat Rev Mol Cell Biol. 2008;9:532–42. - PubMed
-
- Galluzzi L, Morselli E, Kepp O, Vitale I, Rigoni A, Vacchelli E, et al.Mitochondrial gateways to cancer. Molecular Aspects of Medicine. 2010;31:1–20. - PubMed
-
- Radi E, Formichi P, Battisti C, Federico A. Apoptosis and Oxidative Stress in Neurodegenerative Diseases. Journal of Alzheimer’s Disease. 2014;42:S125–S52. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

