Phosphorylation of Pal2 by the protein kinases Kin1 and Kin2 modulates HAC1 mRNA splicing in the unfolded protein response in yeast
- PMID: 34035143
- PMCID: PMC8285061
- DOI: 10.1126/scisignal.aaz4401
Phosphorylation of Pal2 by the protein kinases Kin1 and Kin2 modulates HAC1 mRNA splicing in the unfolded protein response in yeast
Abstract
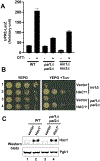
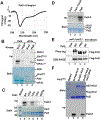
During cellular stress in the budding yeast Saccharomyces cerevisiae, an endoplasmic reticulum (ER)-resident dual kinase and RNase Ire1 splices an intron from HAC1 mRNA in the cytosol, thereby releasing its translational block. Hac1 protein then activates an adaptive cellular stress response called the unfolded protein response (UPR) that maintains ER homeostasis. The polarity-inducing protein kinases Kin1 and Kin2 contribute to HAC1 mRNA processing. Here, we showed that an RNA-protein complex that included the endocytic proteins Pal1 and Pal2 mediated HAC1 mRNA splicing downstream of Kin1 and Kin2. We found that Pal1 and Pal2 bound to the 3' untranslated region (3'UTR) of HAC1 mRNA, and a yeast strain lacking both Pal1 and Pal2 was deficient in HAC1 mRNA processing. We also showed that Kin1 and Kin2 directly phosphorylated Pal2, and that a nonphosphorylatable Pal2 mutant could not rescue the UPR defect in a pal1Δ pal2Δ strain. Thus, our work uncovers a Kin1/2-Pal2 signaling pathway that coordinates HAC1 mRNA processing and ER homeostasis.
Copyright © 2021 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
COMPETING INTERESTS:
The authors declare that they have no competing interests.
Figures






Similar articles
-
A novel role for protein kinase Kin2 in regulating HAC1 mRNA translocation, splicing, and translation.Mol Cell Biol. 2015 Jan;35(1):199-210. doi: 10.1128/MCB.00981-14. Epub 2014 Oct 27. Mol Cell Biol. 2015. PMID: 25348718 Free PMC article.
-
Ribosome depurination by ricin leads to inhibition of endoplasmic reticulum stress-induced HAC1 mRNA splicing on the ribosome.J Biol Chem. 2019 Nov 22;294(47):17848-17862. doi: 10.1074/jbc.RA119.009128. Epub 2019 Oct 17. J Biol Chem. 2019. PMID: 31624149 Free PMC article.
-
Vps34 and TOR Kinases Coordinate HAC1 mRNA Translation in the Presence or Absence of Ire1-Dependent Splicing.Mol Cell Biol. 2021 Jun 23;41(7):e0066220. doi: 10.1128/MCB.00662-20. Epub 2021 Jun 23. Mol Cell Biol. 2021. PMID: 33972394 Free PMC article.
-
Translation Control of HAC1 by Regulation of Splicing in Saccharomyces cerevisiae.Int J Mol Sci. 2019 Jun 12;20(12):2860. doi: 10.3390/ijms20122860. Int J Mol Sci. 2019. PMID: 31212749 Free PMC article. Review.
-
The unfolded protein response: the dawn of a new field.Proc Jpn Acad Ser B Phys Biol Sci. 2015;91(9):469-80. doi: 10.2183/pjab.91.469. Proc Jpn Acad Ser B Phys Biol Sci. 2015. PMID: 26560836 Free PMC article. Review.
Cited by
-
Functional, transcriptomic, and lipidomic studies of the choC gene encoding a phospholipid methyltransferase in Aspergillus fumigatus.Microbiol Spectr. 2024 Jan 11;12(1):e0216823. doi: 10.1128/spectrum.02168-23. Epub 2023 Nov 27. Microbiol Spectr. 2024. PMID: 38009944 Free PMC article.
-
Multi-omics analysis revealed that the protein kinase MoKin1 affected the cellular response to endoplasmic reticulum stress in the rice blast fungus, Magnaporthe oryzae.BMC Genomics. 2024 May 7;25(1):449. doi: 10.1186/s12864-024-10337-8. BMC Genomics. 2024. PMID: 38714914 Free PMC article.
-
Fundamental and Applicative Aspects of the Unfolded Protein Response in Yeasts.J Fungi (Basel). 2023 Oct 5;9(10):989. doi: 10.3390/jof9100989. J Fungi (Basel). 2023. PMID: 37888245 Free PMC article. Review.
-
Endocytic protein Pal1 regulates appressorium formation and is required for full virulence of Magnaporthe oryzae.Mol Plant Pathol. 2022 Jan;23(1):133-147. doi: 10.1111/mpp.13149. Epub 2021 Oct 12. Mol Plant Pathol. 2022. PMID: 34636149 Free PMC article.
-
Pull-down of Biotinylated RNA and Associated Proteins.Bio Protoc. 2022 Feb 20;12(4):e4331. doi: 10.21769/BioProtoc.4331. eCollection 2022 Feb 20. Bio Protoc. 2022. PMID: 35340298 Free PMC article.
References
-
- Walter P, and Ron D (2011) The Unfolded Protein Response: From Stress Pathway to Homeostatic Regulation. Science 334, 1081–1086 - PubMed
-
- Ron D, and Walter P (2007) Signal integration in the endoplasmic reticulum unfolded protein response. Nat Rev Mol Cell Bio 8, 519–529 - PubMed
-
- Cox JS, and Walter P (1996) A novel mechanism for regulating activity of a transcription factor that controls the unfolded protein response. Cell 87, 391–404 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

