Genomic Determinants Encode the Reactivity and Regioselectivity of Flavin-Dependent Halogenases in Bacterial Genomes and Metagenomes
- PMID: 34042468
- PMCID: PMC8269204
- DOI: 10.1128/mSystems.00053-21
Genomic Determinants Encode the Reactivity and Regioselectivity of Flavin-Dependent Halogenases in Bacterial Genomes and Metagenomes
Abstract
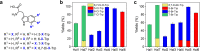
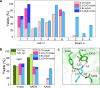
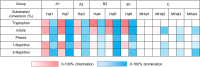
Halogenases create diverse natural products by utilizing halide ions and are of great interest in the synthesis of potential pharmaceuticals and agrochemicals. An increasing number of halogenases discovered in microorganisms are annotated as flavin-dependent halogenases (FDHs), but their chemical reactivities are markedly different and the genomic contents associated with such functional distinction have not been revealed yet. Even though the reactivity and regioselectivity of FDHs are essential in the halogenation activity, these FDHs are annotated inaccurately in the protein sequence repositories without characterizing their functional activities. We carried out a comprehensive sequence analysis and biochemical characterization of FDHs. Using a probabilistic model that we built in this study, FDHs were discovered from 2,787 bacterial genomes and 17 sediment metagenomes. We analyzed the essential genomic determinants that are responsible for substrate binding and subsequent reactions: four flavin adenine dinucleotide-binding, one halide-binding, and four tryptophan-binding sites. Compared with previous studies, our study utilizes large-scale genomic information to propose a comprehensive set of sequence motifs that are related to the active sites and regioselectivity. We reveal that the genomic patterns and phylogenetic locations of the FDHs determine the enzymatic reactivities, which was experimentally validated in terms of the substrate scope and regioselectivity. A large portion of publicly available FDHs needs to be reevaluated to designate their correct functions. Our genomic models establish comprehensive links among genotypic information, reactivity, and regioselectivity of FDHs, thereby laying an important foundation for future discovery and classification of novel FDHs. IMPORTANCE Halogenases are playing an important role as tailoring enzymes in biosynthetic pathways. Flavin-dependent tryptophan halogenases (Trp-FDHs) are among the enzymes that have broad substrate scope and high selectivity. From bacterial genomes and metagenomes, we found highly diverse halogenase sequences by using a well-trained profile hidden Markov model built from the experimentally validated halogenases. The characterization of genotype, steady-state activity, substrate scope, and regioselectivity has established comprehensive links between the information encoded in the genomic sequence and reactivity of FDHs reported here. By constructing models for accurate and detailed sequence markers, our work should guide future discovery and classification of novel FDHs.
Keywords: active sites; genomic patterns; halogenase; mutagenesis; phylogeny; profile hidden Markov model; regioselectivity.
Figures



Similar articles
-
Identifying and Engineering Flavin Dependent Halogenases for Selective Biocatalysis.Acc Chem Res. 2024 Aug 6;57(15):2067-2079. doi: 10.1021/acs.accounts.4c00172. Epub 2024 Jul 22. Acc Chem Res. 2024. PMID: 39038085 Free PMC article.
-
Flavin Adenine Dinucleotide-Dependent Halogenase XanH and Engineering of Multifunctional Fusion Halogenases.Appl Environ Microbiol. 2020 Sep 1;86(18):e01225-20. doi: 10.1128/AEM.01225-20. Print 2020 Sep 1. Appl Environ Microbiol. 2020. PMID: 32651204 Free PMC article.
-
Structures, mechanisms and applications of flavin-dependent halogenases.Enzymes. 2020;47:327-364. doi: 10.1016/bs.enz.2020.05.009. Epub 2020 Jul 18. Enzymes. 2020. PMID: 32951827 Review.
-
A flavin-dependent halogenase from metagenomic analysis prefers bromination over chlorination.PLoS One. 2018 May 10;13(5):e0196797. doi: 10.1371/journal.pone.0196797. eCollection 2018. PLoS One. 2018. PMID: 29746521 Free PMC article.
-
Structural perspective on enzymatic halogenation.Acc Chem Res. 2009 Jan 20;42(1):147-55. doi: 10.1021/ar800088r. Acc Chem Res. 2009. PMID: 18774824 Free PMC article. Review.
Cited by
-
Crystallographic and Thermodynamic Evidence of Negative Coupling in the Flavin-Dependent Tryptophan Halogenases AbeH and BorH.ACS Omega. 2025 Jan 8;10(6):5849-5865. doi: 10.1021/acsomega.4c09590. eCollection 2025 Feb 18. ACS Omega. 2025. PMID: 39989782 Free PMC article.
-
Late-stage diversification of bacterial natural products through biocatalysis.Front Bioeng Biotechnol. 2024 May 14;12:1351583. doi: 10.3389/fbioe.2024.1351583. eCollection 2024. Front Bioeng Biotechnol. 2024. PMID: 38807651 Free PMC article. Review.
References
-
- Gribble GW. 2010. Naturally occurring organohalogen compounds: a comprehensive update. Springer, Verlag, Germany.
-
- Harris CM, Kannan R, Kopecka H, Harris TM. 1985. The role of the chlorine substituents in the antibiotic vancomycin: preparation and characterization of mono- and didechlorovancomycin. J Am Chem Soc 107:6652–6658. doi:10.1021/ja00309a038. - DOI
-
- Rodrigues Pereira E, Belin L, Sancelme M, Prudhomme M, Ollier M, Rapp M, Sevère D, Riou J-F, Fabbro D, Meyer T. 1996. Structure–activity relationships in a series of substituted indolocarbazoles: topoisomerase I and protein kinase C inhibition and antitumoral and antimicrobial properties. J Med Chem 39:4471–4477. doi:10.1021/jm9603779. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

