On the Origin of Frameshift-Robustness of the Standard Genetic Code
- PMID: 34043802
- PMCID: PMC8476161
- DOI: 10.1093/molbev/msab164
On the Origin of Frameshift-Robustness of the Standard Genetic Code
Abstract
The standard genetic code (SGC) has been extensively analyzed for the biological ramifications of its nonrandom structure. For instance, mismatch errors due to point mutation or mistranslation have an overall smaller effect on the amino acid polar requirement under the SGC than under random genetic codes (RGCs). A similar observation was recently made for frameshift errors, prompting the assertion that the SGC has been shaped by natural selection for frameshift-robustness-conservation of certain amino acid properties upon a frameshift mutation or translational frameshift. However, frameshift-robustness confers no benefit because frameshifts usually create premature stop codons that cause nonsense-mediated mRNA decay or production of nonfunctional truncated proteins. We here propose that the frameshift-robustness of the SGC is a byproduct of its mismatch-robustness. Of 564 amino acid properties considered, the SGC exhibits mismatch-robustness in 93-133 properties and frameshift-robustness in 55 properties, respectively, and that the latter is largely a subset of the former. For each of the 564 real and 564 randomly constructed fake properties of amino acids, there is a positive correlation between mismatch-robustness and frameshift-robustness across one million RGCs; this correlation arises because most amino acid changes resulting from a frameshift are also achievable by a mismatch error. Importantly, the SGC does not show significantly higher frameshift-robustness in any of the 55 properties than RGCs of comparable mismatch-robustness. These findings support that the frameshift-robustness of the SGC need not originate through direct selection and can instead be a site effect of its mismatch-robustness.
Keywords: amino acid property; byproduct; mismatch-robustness; mistranslation; mutation; natural selection.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
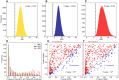
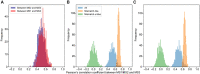
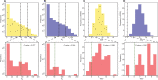
Figures

Similar articles
-
Optimality of circular codes versus the genetic code after frameshift errors.Biosystems. 2020 Jul;195:104134. doi: 10.1016/j.biosystems.2020.104134. Epub 2020 Apr 4. Biosystems. 2020. PMID: 32251681
-
Optimization of the standard genetic code in terms of two mutation types: Point mutations and frameshifts.Biosystems. 2019 Jul;181:44-50. doi: 10.1016/j.biosystems.2019.04.012. Epub 2019 Apr 28. Biosystems. 2019. PMID: 31042561
-
Frameshift and wild-type proteins are often highly similar because the genetic code and genomes were optimized for frameshift tolerance.BMC Genomics. 2022 Jun 2;23(1):416. doi: 10.1186/s12864-022-08435-6. BMC Genomics. 2022. PMID: 35655139 Free PMC article.
-
Origin and Evolution of the Universal Genetic Code.Annu Rev Genet. 2017 Nov 27;51:45-62. doi: 10.1146/annurev-genet-120116-024713. Epub 2017 Aug 30. Annu Rev Genet. 2017. PMID: 28853922 Review.
-
Mathematical fundamentals for the noise immunity of the genetic code.Biosystems. 2018 Feb;164:186-198. doi: 10.1016/j.biosystems.2017.09.007. Epub 2017 Sep 14. Biosystems. 2018. PMID: 28918301 Review.
Cited by
-
Biological factors in the synthetic construction of overlapping genes.BMC Genomics. 2021 Dec 11;22(1):888. doi: 10.1186/s12864-021-08181-1. BMC Genomics. 2021. PMID: 34895142 Free PMC article.
-
Little Evidence the Standard Genetic Code Is Optimized for Resource Conservation.Mol Biol Evol. 2021 Oct 27;38(11):5127-5133. doi: 10.1093/molbev/msab236. Mol Biol Evol. 2021. PMID: 34373928 Free PMC article.
-
Is the Genetic Code Optimized for Resource Conservation?Mol Biol Evol. 2021 Oct 27;38(11):5122-5126. doi: 10.1093/molbev/msab239. Mol Biol Evol. 2021. PMID: 34383940 Free PMC article.
-
Alternative Reading Frames are an Underappreciated Source of Protein Sequence Novelty.J Mol Evol. 2023 Oct;91(5):570-580. doi: 10.1007/s00239-023-10122-3. Epub 2023 Jun 16. J Mol Evol. 2023. PMID: 37326679 Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

