A tri-tuple coordinate system derived for fast and accurate analysis of the colored de Bruijn graph-based pangenomes
- PMID: 34044757
- PMCID: PMC8161984
- DOI: 10.1186/s12859-021-04149-w
A tri-tuple coordinate system derived for fast and accurate analysis of the colored de Bruijn graph-based pangenomes
Abstract
Background: With the rapid development of accurate sequencing and assembly technologies, an increasing number of high-quality chromosome-level and haplotype-resolved assemblies of genomic sequences have been derived, from which there will be great opportunities for computational pangenomics. Although genome graphs are among the most useful models for pangenome representation, their structural complexity makes it difficult to present genome information intuitively, such as the linear reference genome. Thus, efficiently and accurately analyzing the genome graph spatial structure and coordinating the information remains a substantial challenge.
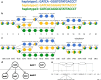
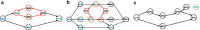
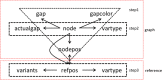
Results: We developed a new method, a colored superbubble (cSupB), that can overcome the complexity of graphs and organize a set of species- or population-specific haplotype sequences of interest. Based on this model, we propose a tri-tuple coordinate system that combines an offset value, topological structure and sample information. Additionally, cSupB provides a novel method that utilizes complete topological information and efficiently detects small indels (< 50 bp) for highly similar samples, which can be validated by simulated datasets. Moreover, we demonstrated that cSupB can adapt to the complex cycle structure.
Conclusions: Although the solution is made suitable for increasingly complex genome graphs by relaxing the constraint, the directed acyclic graph, the motif cSupB and the cSupB method can be extended to any colored directed acyclic graph. We anticipate that our method will facilitate the analysis of individual haplotype variants and population genomic diversity. We have developed a C + + program for implementing our method that is available at https://github.com/eggleader/cSupB .
Keywords: Coordinate system; Genome graph; Variant detection.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
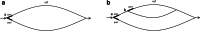
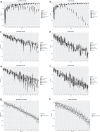

Similar articles
-
Minimizer-space de Bruijn graphs: Whole-genome assembly of long reads in minutes on a personal computer.Cell Syst. 2021 Oct 20;12(10):958-968.e6. doi: 10.1016/j.cels.2021.08.009. Epub 2021 Sep 14. Cell Syst. 2021. PMID: 34525345 Free PMC article.
-
A space and time-efficient index for the compacted colored de Bruijn graph.Bioinformatics. 2018 Jul 1;34(13):i169-i177. doi: 10.1093/bioinformatics/bty292. Bioinformatics. 2018. PMID: 29949982 Free PMC article.
-
Building large updatable colored de Bruijn graphs via merging.Bioinformatics. 2019 Jul 15;35(14):i51-i60. doi: 10.1093/bioinformatics/btz350. Bioinformatics. 2019. PMID: 31510647 Free PMC article.
-
A stepwise guide for pangenome development in crop plants: an alfalfa (Medicago sativa) case study.BMC Genomics. 2024 Oct 31;25(1):1022. doi: 10.1186/s12864-024-10931-w. BMC Genomics. 2024. PMID: 39482604 Free PMC article. Review.
-
A survey of sequence-to-graph mapping algorithms in the pangenome era.Genome Biol. 2025 May 22;26(1):138. doi: 10.1186/s13059-025-03606-6. Genome Biol. 2025. PMID: 40405275 Free PMC article. Review.
Cited by
-
ploidyfrost: Reference-free estimation of ploidy level from whole genome sequencing data based on de Bruijn graphs.Mol Ecol Resour. 2023 Feb;23(2):499-510. doi: 10.1111/1755-0998.13720. Epub 2022 Nov 1. Mol Ecol Resour. 2023. PMID: 36239149 Free PMC article.
-
A gentle introduction to pangenomics.Brief Bioinform. 2024 Sep 23;25(6):bbae588. doi: 10.1093/bib/bbae588. Brief Bioinform. 2024. PMID: 39552065 Free PMC article. Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

