Calcium-Collagen Coupling is Vital for Biomineralization Schedule
- PMID: 34047068
- PMCID: PMC8336496
- DOI: 10.1002/advs.202100363
Calcium-Collagen Coupling is Vital for Biomineralization Schedule
Abstract
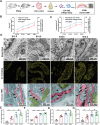
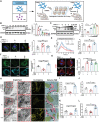
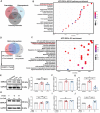
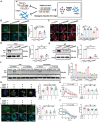
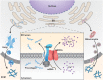
Biomineralization is a chemical reaction that occurs in organisms in which collagen initiates and guides the growth and crystallization of matched apatite minerals. However, there is little known about the demand pattern for calcium salts and collagen needed by biomineralization. In this study, natural bone biomineralization is analyzed, and a novel interplay between calcium concentration and collagen production is observed. Any quantitative change in one of the entities causes a corresponding change in the other. Translocation-associated membrane protein 2 (TRAM2) is identified as an intermediate factor whose silencing disrupts this relationship and causes poor mineralization. TRAM2 directly interacts with the sarcoplasmic/endoplasmic reticulum calcium ATPase 2b (SERCA2b) and modulates SERCA2b activity to couple calcium enrichment with collagen biosynthesis. Collectively, these findings indicate that osteoblasts can independently and directly regulate the process of biomineralization via this coupling. This knowledge has significant implications for the developmentally inspired design of biomaterials for bone regenerative applications.
Keywords: biomineralization; bone biomaterials; bones; calcium-collagen coupling; collagen.
© 2021 The Authors. Advanced Science published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Enzymatic Approach in Calcium Phosphate Biomineralization: A Contribution to Reconcile the Physicochemical with the Physiological View.Int J Mol Sci. 2021 Nov 30;22(23):12957. doi: 10.3390/ijms222312957. Int J Mol Sci. 2021. PMID: 34884758 Free PMC article. Review.
-
TRAM2 protein interacts with endoplasmic reticulum Ca2+ pump Serca2b and is necessary for collagen type I synthesis.Mol Cell Biol. 2004 Feb;24(4):1758-68. doi: 10.1128/MCB.24.4.1758-1768.2004. Mol Cell Biol. 2004. PMID: 14749390 Free PMC article.
-
Phosphatidylserine controls calcium phosphate nucleation and growth on lipid monolayers: A physicochemical understanding of matrix vesicle-driven biomineralization.J Struct Biol. 2020 Nov 1;212(2):107607. doi: 10.1016/j.jsb.2020.107607. Epub 2020 Aug 26. J Struct Biol. 2020. PMID: 32858148 Free PMC article.
-
First evidence of octacalcium phosphate@osteocalcin nanocomplex as skeletal bone component directing collagen triple-helix nanofibril mineralization.Sci Rep. 2018 Sep 12;8(1):13696. doi: 10.1038/s41598-018-31983-5. Sci Rep. 2018. PMID: 30209287 Free PMC article.
-
Hormonal regulation of biomineralization.Nat Rev Endocrinol. 2021 May;17(5):261-275. doi: 10.1038/s41574-021-00477-2. Epub 2021 Mar 16. Nat Rev Endocrinol. 2021. PMID: 33727709 Review.
Cited by
-
High-Performance Hydrogel-Encapsulated Engineered Exosomes for Supporting Endoplasmic Reticulum Homeostasis and Boosting Diabetic Bone Regeneration.Adv Sci (Weinh). 2024 May;11(17):e2309491. doi: 10.1002/advs.202309491. Epub 2024 Feb 21. Adv Sci (Weinh). 2024. PMID: 38380490 Free PMC article.
-
Advances in Engineered Three-Dimensional (3D) Body Articulation Unit Models.Drug Des Devel Ther. 2022 Jan 18;16:213-235. doi: 10.2147/DDDT.S344036. eCollection 2022. Drug Des Devel Ther. 2022. PMID: 35087267 Free PMC article. Review.
-
Phosphorylated and Phosphonated Low-Complexity Protein Segments for Biomimetic Mineralization and Repair of Tooth Enamel.Adv Sci (Weinh). 2022 Feb;9(6):e2103829. doi: 10.1002/advs.202103829. Epub 2022 Jan 2. Adv Sci (Weinh). 2022. PMID: 34978158 Free PMC article.
-
Enzymatic Approach in Calcium Phosphate Biomineralization: A Contribution to Reconcile the Physicochemical with the Physiological View.Int J Mol Sci. 2021 Nov 30;22(23):12957. doi: 10.3390/ijms222312957. Int J Mol Sci. 2021. PMID: 34884758 Free PMC article. Review.
-
Collagen structures of demineralized bone paper direct mineral metabolism.JBMR Plus. 2024 Jun 17;8(8):ziae080. doi: 10.1093/jbmrpl/ziae080. eCollection 2024 Aug. JBMR Plus. 2024. PMID: 38989259 Free PMC article.
References
-
- a) Wang S. J., Jiang D., Zhang Z. Z., Chen Y. R., Yang Z. D., Zhang J. Y., Shi J., Wang X., Yu J. K., Adv. Mater. 2019, 31, 1904341; - PubMed
- b) Yuan X., Wei Y., Villasante A., Ng J. J. D., Arkonac D. E., Chao P. G., Vunjak‐Novakovic G., Biomaterials 2017, 132, 59; - PMC - PubMed
- c) Lee C. S., Kim S., Fan J., Hwang H. S., Aghaloo T., Lee M., Sci. Adv. 2020, 6, eaaz7822; - PMC - PubMed
- d) McDermott A. M., Herberg S., Mason D. E., Collins J. M., Pearson H. B., Dawahare J. H., Tang R., Patwa A. N., Grinstaff M. W., Kelly D. J., Alsberg E., Boerckel J. D., Sci. Transl. Med. 2019, 11, eaav7756. - PMC - PubMed
-
- Wang Y., Azais T., Robin M., Vallee A., Catania C., Legriel P., Pehau‐Arnaudet G., Babonneau F., Giraud‐Guille M. M., Nassif N., Nat. Mater. 2012, 11, 724. - PubMed
-
- a) Kim D., Lee B., Thomopoulos S., Jun Y. S., Nat. Commun. 2018, 9, 962; - PMC - PubMed
- b) Song Q., Jiao K., Tonggu L., Wang L. G., Zhang S. L., Yang Y. D., Zhang L., Bian J. H., Hao D. X., Wang C. Y., Ma Y. X., Arola D. D., Breschi L., Chen J. H., Tay F. R., Niu L. N., Sci. Adv. 2019, 5, eaav9075. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
