A PorX/PorY and σP Feedforward Regulatory Loop Controls Gene Expression Essential for Porphyromonas gingivalis Virulence
- PMID: 34047648
- PMCID: PMC8265659
- DOI: 10.1128/mSphere.00428-21
A PorX/PorY and σP Feedforward Regulatory Loop Controls Gene Expression Essential for Porphyromonas gingivalis Virulence
Abstract
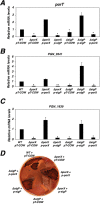
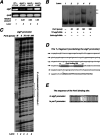
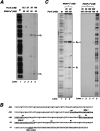
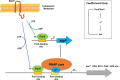
The PorX/PorY two-component system in the periodontal pathogen Porphyromonas gingivalis controls the expression of the por genes, encoding a type IX secretion system, and the sigP gene, encoding sigma factor σP. Previous results implied that PorX/PorY and σP formed a regulatory cascade because the PorX/PorY-activated σP enhanced the por genes, including porT, via binding to their promoters. We recently showed that PorX also binds to the por promoters, thus suggesting that an alternative mechanism is required for the PorX/PorY- and σP-governed expression. Here, our in vitro assays show the PorX response regulator binds to the sigP promoter at a sequence shared with the porT promoter and enhances its transcription, mediated by a reconstituted P. gingivalis RNA polymerase holoenzyme. Merely producing σP in trans fails to reverse the porT transcription in a porX mutant, which further argues against the action of the proposed regulatory cascade. An in vitro transcription assay using a reconstituted RNA polymerase-σP holoenzyme verifies the direct role of PorX in porT transcription, since transcription is enhanced by a pure PorX protein. Accordingly, we propose that the PorX/PorY system coordinates with σP to construct a coherent regulatory mechanism, known as the feedforward loop. Specifically, PorX will not only bind to the sigP promoter to stimulate the expression of σP, but also bind to the porT promoter to facilitate the RNA polymerase-σP-dependent transcription. Importantly, mutations at the porX and sigP genes attenuate bacterial virulence in a mouse model, demonstrating that this regulatory mechanism is essential for P. gingivalis pathogenesis. IMPORTANCE The anaerobic bacterium Porphyromonas gingivalis is not only the major etiologic agent for chronic periodontitis, but also prevalent in some common noncommunicable diseases such as cardiovascular disease, Alzheimer's disease, and rheumatoid arthritis. We present genetic, biochemical, and biological results to demonstrate that the PorX/PorY two-component system and sigma factor σP build a specific regulatory network to coordinately control transcription of the genes encoding the type IX secretion system, and perhaps also other virulence factors. Results in this study verify that the response regulator PorX stimulates the expression of the genes encoding both σP and the type IX secretion system by binding to their promoters. This study also provides evidence that σP, like the PorX/PorY system, contributes to P. gingivalis virulence in a mouse model.
Keywords: PorX/PorY; Porphyromonas gingivalis; extracytoplasmic function sigma factor P; feedforward loop; in vitro transcription; transcription regulation; two-component system; type IX secretion system; virulence factors.
Figures


Similar articles
-
The PorX/PorY system is a virulence factor of Porphyromonas gingivalis and mediates the activation of the type IX secretion system.J Biol Chem. 2021 Jan-Jun;296:100574. doi: 10.1016/j.jbc.2021.100574. Epub 2021 Mar 21. J Biol Chem. 2021. PMID: 33757767 Free PMC article.
-
A two-component system regulates gene expression of the type IX secretion component proteins via an ECF sigma factor.Sci Rep. 2016 Mar 21;6:23288. doi: 10.1038/srep23288. Sci Rep. 2016. PMID: 26996145 Free PMC article.
-
The PorX Response Regulator of the Porphyromonas gingivalis PorXY Two-Component System Does Not Directly Regulate the Type IX Secretion Genes but Binds the PorL Subunit.Front Cell Infect Microbiol. 2016 Aug 31;6:96. doi: 10.3389/fcimb.2016.00096. eCollection 2016. Front Cell Infect Microbiol. 2016. PMID: 27630829 Free PMC article.
-
Molecular Strategies Underlying Porphyromonas gingivalis Virulence.J Mol Biol. 2021 Apr 2;433(7):166836. doi: 10.1016/j.jmb.2021.166836. Epub 2021 Feb 1. J Mol Biol. 2021. PMID: 33539891 Review.
-
The Type IX Secretion System: Advances in Structure, Function and Organisation.Microorganisms. 2020 Aug 1;8(8):1173. doi: 10.3390/microorganisms8081173. Microorganisms. 2020. PMID: 32752268 Free PMC article. Review.
Cited by
-
Response regulator PorX coordinates oligonucleotide signalling and gene expression to control the secretion of virulence factors.Nucleic Acids Res. 2022 Nov 28;50(21):12558-12577. doi: 10.1093/nar/gkac1103. Nucleic Acids Res. 2022. PMID: 36464236 Free PMC article.
-
Hemin availability induces coordinated DNA methylation and gene expression changes in Porphyromonas gingivalis.mSystems. 2023 Aug 31;8(4):e0119322. doi: 10.1128/msystems.01193-22. Epub 2023 Jul 12. mSystems. 2023. PMID: 37436062 Free PMC article.
-
PorA of the Type IX Secretion Is a Ligand of the PorXY Two-Component Regulatory System in Porphyromonas gingivalis.Mol Microbiol. 2025 Jun;123(6):569-585. doi: 10.1111/mmi.15363. Epub 2025 Apr 8. Mol Microbiol. 2025. PMID: 40195800 Free PMC article.
-
Unveiling the molecular mechanisms of the type IX secretion system's response regulator: Structural and functional insights.PNAS Nexus. 2024 Jul 31;3(8):pgae316. doi: 10.1093/pnasnexus/pgae316. eCollection 2024 Aug. PNAS Nexus. 2024. PMID: 39139265 Free PMC article.
-
Purification, crystallization and crystallographic analysis of the PorX response regulator associated with the type IX secretion system.Acta Crystallogr F Struct Biol Commun. 2022 Oct 1;78(Pt 10):354-362. doi: 10.1107/S2053230X22008500. Epub 2022 Sep 26. Acta Crystallogr F Struct Biol Commun. 2022. PMID: 36189719 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
