Frequent loss of heterozygosity in CRISPR-Cas9-edited early human embryos
- PMID: 34050011
- PMCID: PMC8179174
- DOI: 10.1073/pnas.2004832117
Frequent loss of heterozygosity in CRISPR-Cas9-edited early human embryos
Abstract
CRISPR-Cas9 genome editing is a promising technique for clinical applications, such as the correction of disease-associated alleles in somatic cells. The use of this approach has also been discussed in the context of heritable editing of the human germ line. However, studies assessing gene correction in early human embryos report low efficiency of mutation repair, high rates of mosaicism, and the possibility of unintended editing outcomes that may have pathologic consequences. We developed computational pipelines to assess single-cell genomics and transcriptomics datasets from OCT4 (POU5F1) CRISPR-Cas9-targeted and control human preimplantation embryos. This allowed us to evaluate on-target mutations that would be missed by more conventional genotyping techniques. We observed loss of heterozygosity in edited cells that spanned regions beyond the POU5F1 on-target locus, as well as segmental loss and gain of chromosome 6, on which the POU5F1 gene is located. Unintended genome editing outcomes were present in ∼16% of the human embryo cells analyzed and spanned 4-20 kb. Our observations are consistent with recent findings indicating complexity at on-target sites following CRISPR-Cas9 genome editing. Our work underscores the importance of further basic research to assess the safety of genome editing techniques in human embryos, which will inform debates about the potential clinical use of this technology.
Keywords: CRISPR-Cas9; genome editing; human embryo; loss of heterozygosity; segmental aneuploidy.
Conflict of interest statement
The authors declare no competing interest.
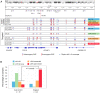
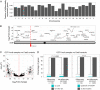
Figures


Similar articles
-
Comparative analysis of mouse and human preimplantation development following POU5F1 CRISPR/Cas9 targeting reveals interspecies differences.Hum Reprod. 2021 Apr 20;36(5):1242-1252. doi: 10.1093/humrep/deab027. Hum Reprod. 2021. PMID: 33609360
-
TEAD4 regulates trophectoderm differentiation upstream of CDX2 in a GATA3-independent manner in the human preimplantation embryo.Hum Reprod. 2022 Jul 30;37(8):1760-1773. doi: 10.1093/humrep/deac138. Hum Reprod. 2022. PMID: 35700449
-
Genome editing reveals a role for OCT4 in human embryogenesis.Nature. 2017 Oct 5;550(7674):67-73. doi: 10.1038/nature24033. Epub 2017 Sep 20. Nature. 2017. PMID: 28953884 Free PMC article.
-
Mosaicism in CRISPR/Cas9-mediated genome editing.Dev Biol. 2019 Jan 15;445(2):156-162. doi: 10.1016/j.ydbio.2018.10.008. Epub 2018 Oct 22. Dev Biol. 2019. PMID: 30359560 Review.
-
Genome engineering through CRISPR/Cas9 technology in the human germline and pluripotent stem cells.Hum Reprod Update. 2016 Jun;22(4):411-9. doi: 10.1093/humupd/dmw005. Epub 2016 Feb 29. Hum Reprod Update. 2016. PMID: 26932460 Review.
Cited by
-
CRISPR-Cas and Its Wide-Ranging Applications: From Human Genome Editing to Environmental Implications, Technical Limitations, Hazards and Bioethical Issues.Cells. 2021 Apr 21;10(5):969. doi: 10.3390/cells10050969. Cells. 2021. PMID: 33919194 Free PMC article. Review.
-
CRISPR, animals, and FDA oversight: Building a path to success.Proc Natl Acad Sci U S A. 2021 Jun 1;118(22):e2004831117. doi: 10.1073/pnas.2004831117. Epub 2021 Apr 30. Proc Natl Acad Sci U S A. 2021. PMID: 34050010 Free PMC article.
-
Multi-pathway DNA-repair reporters reveal competition between end-joining, single-strand annealing and homologous recombination at Cas9-induced DNA double-strand breaks.Nat Commun. 2022 Sep 8;13(1):5295. doi: 10.1038/s41467-022-32743-w. Nat Commun. 2022. PMID: 36075911 Free PMC article.
-
Detection of unintended on-target effects in CRISPR genome editing by DNA donors carrying diagnostic substitutions.Nucleic Acids Res. 2023 Mar 21;51(5):e26. doi: 10.1093/nar/gkac1254. Nucleic Acids Res. 2023. PMID: 36620901 Free PMC article.
-
Methods and applications for single-cell and spatial multi-omics.Nat Rev Genet. 2023 Aug;24(8):494-515. doi: 10.1038/s41576-023-00580-2. Epub 2023 Mar 2. Nat Rev Genet. 2023. PMID: 36864178 Free PMC article. Review.
References
-
- A Lea R., K Niakan K., Human germline genome editing. Nat. Cell Biol. 21, 1479–1489 (2019). - PubMed
-
- National Academy of Medicine, National Academy of Sciences, The Royal Society , Heritable Human Genome Editing (The National Academies Press, 2020). - PubMed
-
- Nuffield Council on Bioethics , Genome Editing and Human Reproduction: Social and Ethical Issues (Nuffield Council on Bioethics, 2018).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

