DNA polymerase III protein, HolC, helps resolve replication/transcription conflicts
- PMID: 34055967
- PMCID: PMC8144910
- DOI: 10.15698/mic2021.06.753
DNA polymerase III protein, HolC, helps resolve replication/transcription conflicts
Abstract
In Escherichia coli, DNA replication is catalyzed by an assembly of proteins, the DNA polymerase III holoenzyme. This complex includes the polymerase and proofreading subunits, the processivity clamp and clamp loader complex. The holC gene encodes an accessory protein (known as χ) to the core clamp loader complex and is the only protein of the holoenzyme that binds to single-strand DNA binding protein, SSB. HolC is not essential for viability although mutants show growth impairment, genetic instability and sensitivity to DNA damaging agents. In this study we isolate spontaneous suppressor mutants in a holCΔ strain and identify these by whole genome sequencing. Some suppressors are alleles of RNA polymerase, suggesting that transcription is problematic for holC mutant strains, and of sspA, stringent starvation protein. Using a conditional holC plasmid, we examine factors affecting transcription elongation and termination for synergistic or suppressive effects on holC mutant phenotypes. Alleles of RpoA (α), RpoB (β) and RpoC (β') RNA polymerase holoenzyme can partially suppress loss of HolC. In contrast, mutations in transcription factors DksA and NusA enhanced the inviability of holC mutants. HolC mutants showed enhanced sensitivity to bicyclomycin, a specific inhibitor of Rho-dependent termination. Bicyclomycin also reverses suppression of holC by rpoA, rpoC and sspA. An inversion of the highly expressed rrnA operon exacerbates the growth defects of holC mutants. We propose that transcription complexes block replication in holC mutants and Rho-dependent transcriptional termination and DksA function are particularly important to sustain viability and chromosome integrity.
Keywords: DNA polymerase; RNA polymerase; Rho; transcription termination.
Copyright: © 2021 Lovett.
Conflict of interest statement
Conflict of Interest: The author declares no conflict of interest.
Figures
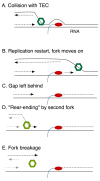
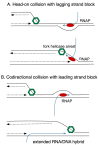
Comment on
-
The Role of Replication Clamp-Loader Protein HolC of Escherichia coli in Overcoming Replication/Transcription Conflicts.mBio. 2021 Mar 9;12(2):e00184-21. doi: 10.1128/mBio.00184-21. mBio. 2021. PMID: 33688004 Free PMC article.
Similar articles
-
The Role of Replication Clamp-Loader Protein HolC of Escherichia coli in Overcoming Replication/Transcription Conflicts.mBio. 2021 Mar 9;12(2):e00184-21. doi: 10.1128/mBio.00184-21. mBio. 2021. PMID: 33688004 Free PMC article.
-
Alternative complexes formed by the Escherichia coli clamp loader accessory protein HolC (x) with replication protein HolD (ψ) and repair protein YoaA.DNA Repair (Amst). 2021 Apr;100:103006. doi: 10.1016/j.dnarep.2020.103006. Epub 2021 Feb 2. DNA Repair (Amst). 2021. PMID: 33582602 Free PMC article.
-
A Screen for rfaH Suppressors Reveals a Key Role for a Connector Region of Termination Factor Rho.mBio. 2017 May 30;8(3):e00753-17. doi: 10.1128/mBio.00753-17. mBio. 2017. PMID: 28559482 Free PMC article.
-
Autogenous and post-transcriptional regulation of RNA polymerase synthesis.Mol Cell Biochem. 1980 Aug 16;31(3):177-96. doi: 10.1007/BF00225850. Mol Cell Biochem. 1980. PMID: 7003354 Review.
-
DksA and DNA double-strand break repair.Curr Genet. 2019 Dec;65(6):1297-1300. doi: 10.1007/s00294-019-00983-x. Epub 2019 May 10. Curr Genet. 2019. PMID: 31076845 Review.
Cited by
-
Comparative genomics reveals intraspecific divergence of Acidithiobacillus ferrooxidans: insights from evolutionary adaptation.Microb Genom. 2023 Jun;9(6):mgen001038. doi: 10.1099/mgen.0.001038. Microb Genom. 2023. PMID: 37285209 Free PMC article.
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
