Impacts of allopolyploidization and structural variation on intraspecific diversification in Brassica rapa
- PMID: 34059118
- PMCID: PMC8166115
- DOI: 10.1186/s13059-021-02383-2
Impacts of allopolyploidization and structural variation on intraspecific diversification in Brassica rapa
Abstract
Background: Despite the prevalence and recurrence of polyploidization in the speciation of flowering plants, its impacts on crop intraspecific genome diversification are largely unknown. Brassica rapa is a mesopolyploid species that is domesticated into many subspecies with distinctive morphotypes.
Results: Herein, we report the consequences of the whole-genome triplication (WGT) on intraspecific diversification using a pan-genome analysis of 16 de novo assembled and two reported genomes. Among the genes that derive from WGT, 13.42% of polyploidy-derived genes accumulate more transposable elements and non-synonymous mutations than other genes during individual genome evolution. We denote such genes as being "flexible." We construct the Brassica rapa ancestral genome and observe the continuing influence of the dominant subgenome on intraspecific diversification in B. rapa. The gene flexibility is biased to the more fractionated subgenomes (MFs), in contrast to the more intact gene content of the dominant LF (least fractionated) subgenome. Furthermore, polyploidy-derived flexible syntenic genes are implicated in the response to stimulus and the phytohormone auxin; this may reflect adaptation to the environment. Using an integrated graph-based genome, we investigate the structural variation (SV) landscapes in 524 B. rapa genomes. We observe that SVs track morphotype domestication. Four out of 266 candidate genes for Chinese cabbage domestication are speculated to be involved in the leafy head formation.
Conclusions: This pan-genome uncovers the possible contributions of allopolyploidization on intraspecific diversification and the possible and underexplored role of SVs in favorable trait domestication. Collectively, our work serves as a rich resource for genome-based B. rapa improvement.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




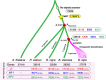

References
-
- Tank DC, Eastman JM, Pennell MW, Soltis PS, Soltis DE, Hinchliff CE, Brown JW, Sessa EB, Harmon LJ. Nested radiations and the pulse of angiosperm diversification: increased diversification rates often follow whole genome duplications. New Phytol. 2015;207(2):454–467. doi: 10.1111/nph.13491. - DOI - PubMed
-
- Zhang K, Wang XW, Cheng F. Plant polyploidy: origin, evolution, and its influence on crop domestication. Horticultural Plant J. 2019;5:231–239. doi: 10.1016/j.hpj.2019.11.003. - DOI
Publication types
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources

