A Protocol for Untargeted Metabolomic Analysis: From Sample Preparation to Data Processing
- PMID: 34060055
- PMCID: PMC9284939
- DOI: 10.1007/978-1-0716-1266-8_27
A Protocol for Untargeted Metabolomic Analysis: From Sample Preparation to Data Processing
Abstract
Untargeted metabolomics has rapidly become a profiling method of choice in many areas of research, including mitochondrial biology. Most commonly, untargeted metabolomics is performed with liquid chromatography/mass spectrometry because it enables measurement of a relatively wide range of physiochemically diverse molecules. Specifically, to assess energy pathways that are associated with mitochondrial metabolism, hydrophilic interaction liquid chromatography (HILIC) is often applied before analysis with a high-resolution accurate mass instrument. The workflow produces large, complex data files that are impractical to analyze manually. Here, we present a protocol to perform untargeted metabolomics on biofluids such as plasma, urine, and cerebral spinal fluid with a HILIC separation and an Orbitrap mass spectrometer. Our protocol describes each step of the analysis in detail, from preparation of solvents for chromatography to selecting parameters during data processing.
Keywords: Accurate mass; Data-dependent acquisition; HILIC; High-resolution; Liquid chromatography; Mass spectrometry; Metabolites; Metabolomics; Profiling; Quality assurance; Quality control.
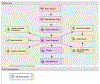
Figures



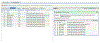


Similar articles
-
Chromatographic Choice in Cerebrospinal Fluid Metabolomics Identification Outcome : Metabolomics for Cerebrospinal Fluid.Methods Mol Biol. 2025;2925:91-101. doi: 10.1007/978-1-0716-4534-5_5. Methods Mol Biol. 2025. PMID: 40498182
-
An exploratory approach for an oriented development of an untargeted hydrophilic interaction liquid chromatography-mass spectrometry platform for polar metabolites in biological matrices.J Chromatogr A. 2021 Jan 25;1637:461807. doi: 10.1016/j.chroma.2020.461807. Epub 2020 Dec 15. J Chromatogr A. 2021. PMID: 33360078
-
Evaluation of the technical variations and the suitability of a hydrophilic interaction liquid chromatography-high resolution mass spectrometry (ZIC-pHILIC-Exactive orbitrap) for clinical urinary metabolomics study.J Chromatogr B Analyt Technol Biomed Life Sci. 2016 Jun 1;1022:199-205. doi: 10.1016/j.jchromb.2016.04.017. Epub 2016 Apr 13. J Chromatogr B Analyt Technol Biomed Life Sci. 2016. PMID: 27107246
-
Metabolomic applications of HILIC-LC-MS.Mass Spectrom Rev. 2010 Sep-Oct;29(5):671-84. doi: 10.1002/mas.20252. Mass Spectrom Rev. 2010. PMID: 19557839 Review.
-
Mass spectrometric based approaches in urine metabolomics and biomarker discovery.Mass Spectrom Rev. 2017 Mar;36(2):115-134. doi: 10.1002/mas.21455. Epub 2015 Apr 16. Mass Spectrom Rev. 2017. PMID: 25881008 Review.
Cited by
-
Immunogenic shift of arginine metabolism triggers systemic metabolic and immunological reprogramming to prevent HER2+ breast cancer.bioRxiv [Preprint]. 2024 Oct 26:2024.10.23.619827. doi: 10.1101/2024.10.23.619827. bioRxiv. 2024. Update in: Cancer Metab. 2025 Mar 20;13(1):15. doi: 10.1186/s40170-025-00384-4. PMID: 39484369 Free PMC article. Updated. Preprint.
-
Effects of Dietary Medium-Chain Triglyceride Supplementation on the Serum Metabolome of Young Adult and Senior Canines.Animals (Basel). 2024 Dec 11;14(24):3577. doi: 10.3390/ani14243577. Animals (Basel). 2024. PMID: 39765481 Free PMC article.
-
Precision Dietary Intervention: Gut Microbiome and Meta-metabolome as Functional Readouts.Phenomics. 2025 Feb 14;5(1):23-50. doi: 10.1007/s43657-024-00193-7. eCollection 2025 Feb. Phenomics. 2025. PMID: 40313608 Review.
-
Plant Growth Promotion and Plant Disease Suppression Induced by Bacillus amyloliquefaciens Strain GD4a.Plants (Basel). 2024 Feb 28;13(5):672. doi: 10.3390/plants13050672. Plants (Basel). 2024. PMID: 38475518 Free PMC article.
-
Global metabolomic profiling of tumor tissue and paired serum samples to identify biomarkers for response to neoadjuvant FOLFIRINOX treatment of human pancreatic cancer.Mol Oncol. 2025 Feb;19(2):391-411. doi: 10.1002/1878-0261.13759. Epub 2024 Nov 15. Mol Oncol. 2025. PMID: 39545923 Free PMC article.
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

