Non-Random Genome Editing and Natural Cellular Engineering in Cognition-Based Evolution
- PMID: 34066959
- PMCID: PMC8148535
- DOI: 10.3390/cells10051125
Non-Random Genome Editing and Natural Cellular Engineering in Cognition-Based Evolution
Abstract
Neo-Darwinism presumes that biological variation is a product of random genetic replication errors and natural selection. Cognition-Based Evolution (CBE) asserts a comprehensive alternative approach to phenotypic variation and the generation of biological novelty. In CBE, evolutionary variation is the product of natural cellular engineering that permits purposive genetic adjustments as cellular problem-solving. CBE upholds that the cornerstone of biology is the intelligent measuring cell. Since all biological information that is available to cells is ambiguous, multicellularity arises from the cellular requirement to maximize the validity of available environmental information. This is best accomplished through collective measurement purposed towards maintaining and optimizing individual cellular states of homeorhesis as dynamic flux that sustains cellular equipoise. The collective action of the multicellular measurement and assessment of information and its collaborative communication is natural cellular engineering. Its yield is linked cellular ecologies and mutualized niche constructions that comprise biofilms and holobionts. In this context, biological variation is the product of collective differential assessment of ambiguous environmental cues by networking intelligent cells. Such concerted action is enabled by non-random natural genomic editing in response to epigenetic impacts and environmental stresses. Random genetic activity can be either constrained or deployed as a 'harnessing of stochasticity'. Therefore, genes are cellular tools. Selection filters cellular solutions to environmental stresses to assure continuous cellular-organismal-environmental complementarity. Since all multicellular eukaryotes are holobionts as vast assemblages of participants of each of the three cellular domains (Prokaryota, Archaea, Eukaryota) and the virome, multicellular variation is necessarily a product of co-engineering among them.
Keywords: cognition-based evolution; holobiont; natural cellular engineering; natural genetic engineering; niche construction; self-reference; senome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

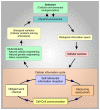
References
-
- Koonin E.V. The Logic of Chance: The Nature and Origin of Biological Evolution. Pearson Education; Upper Saddle River, NJ, USA: 2012.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

