Comparative Gene Signature of (-)-Oleocanthal Formulation Treatments in Heterogeneous Triple Negative Breast Tumor Models: Oncological Therapeutic Target Insights
- PMID: 34069906
- PMCID: PMC8157589
- DOI: 10.3390/nu13051706
Comparative Gene Signature of (-)-Oleocanthal Formulation Treatments in Heterogeneous Triple Negative Breast Tumor Models: Oncological Therapeutic Target Insights
Abstract
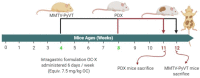
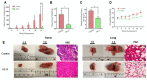
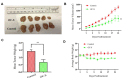
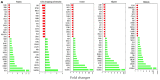
Triple negative breast cancer (TNBC) heterogeneity and limited therapeutic options confer its phenotypic aggressiveness. The discovery of anti-TNBC natural products with valid molecular target(s) and defined pharmacodynamic profile would facilitate their therapeutic nutraceutical use by TNBC patients. The extra-virgin olive oil (EVOO) is a key Mediterranean diet ingredient. S-(-)-Oleocanthal (OC) leads the bioactive anti-tumor EVOO phenolic ingredients. A previous study reported the solid dispersion formulated OC with (+)-xylitol (OC-X) suppressed the in vivo progression and recurrence of the TNBC MDA-MB-231 cells. This study investigates the ability of OC-X formulation to suppress the in vivo heterogeneous BC initiation and progression utilizing advanced preclinical transgenic MMTV-PyVT and TNBC PDX mouse models. Furthermore, the clustering of the gene expression profiles in MMTV-PyVT and PDX mouse tumors treated with OC-X acquired by a Clariom S microarray analysis identified the distinctly affected genes. Several affected novel signature genes identified in response to OC-X treatments and proved overlapped in both mouse and human tumor models, shedding some lights toward understanding the OC anticancer molecular mechanism and assisting in predicting prospective clinical outcomes. This study provides molecular and preclinical evidences of OC-X potential as a nutraceutical suppressing heterogeneous TNBC model and offers preliminary gene-level therapeutic mechanistic insights.
Keywords: (−)-oleocanthal; MMTV-PyVT; PDX; TNBC; extra-virgin olive oil; microarray.
Conflict of interest statement
K. A. El Sayed is a Medicinal Chemistry Chief Scientific Officer without compensation in the Shreveport, Louisiana-based Oleolive.
Figures



Similar articles
-
Harnessing Nature's Chemistry: Deciphering Olive Oil Phenolics for the Control of Invasive Breast Carcinoma.Molecules. 2025 Jul 28;30(15):3157. doi: 10.3390/molecules30153157. Molecules. 2025. PMID: 40807332 Free PMC article.
-
Novel olive oil phenolic (-)-oleocanthal (+)-xylitol-based solid dispersion formulations with potent oral anti-breast cancer activities.Int J Pharm. 2019 Oct 5;569:118596. doi: 10.1016/j.ijpharm.2019.118596. Epub 2019 Aug 5. Int J Pharm. 2019. PMID: 31394181 Free PMC article.
-
(-)-Oleocanthal as a Dual c-MET-COX2 Inhibitor for the Control of Lung Cancer.Nutrients. 2020 Jun 11;12(6):1749. doi: 10.3390/nu12061749. Nutrients. 2020. PMID: 32545325 Free PMC article.
-
Health-promoting properties of oleocanthal and oleacein: Two secoiridoids from extra-virgin olive oil.Crit Rev Food Sci Nutr. 2020;60(15):2532-2548. doi: 10.1080/10408398.2019.1650715. Epub 2019 Aug 19. Crit Rev Food Sci Nutr. 2020. PMID: 31423808 Review.
-
Extra Virgin Olive Oil Phenolic Compounds: Modulating Mitochondrial Function and Protecting Against Chronic Diseases-A Narrative Review.Nutrients. 2025 Apr 25;17(9):1443. doi: 10.3390/nu17091443. Nutrients. 2025. PMID: 40362752 Free PMC article. Review.
Cited by
-
The Olive Oil Monophenolic Secoiridoid Ligstroside Aglycone Suppresses Melanoma Progression by Targeting the BRAF Signaling Pathway.Molecules. 2025 Jan 1;30(1):139. doi: 10.3390/molecules30010139. Molecules. 2025. PMID: 39795195 Free PMC article.
-
Next-generation sequencing reveals altered gene expression and enriched pathways in triple-negative breast cancer cells treated with oleuropein and oleocanthal.Funct Integr Genomics. 2023 Sep 14;23(4):299. doi: 10.1007/s10142-023-01230-w. Funct Integr Genomics. 2023. PMID: 37707691 Free PMC article.
-
Protective Effects and Benefits of Olive Oil and Its Extracts on Women's Health.Nutrients. 2021 Nov 27;13(12):4279. doi: 10.3390/nu13124279. Nutrients. 2021. PMID: 34959830 Free PMC article. Review.
-
Anticancer Effects of Secoiridoids-A Scoping Review of the Molecular Mechanisms behind the Chemopreventive Effects of the Olive Tree Components Oleocanthal, Oleacein, and Oleuropein.Nutrients. 2024 Aug 18;16(16):2755. doi: 10.3390/nu16162755. Nutrients. 2024. PMID: 39203892 Free PMC article.
-
The Cancer-Associated Antigens Sialyl Lewisa/x and Sda: Two Opposite Faces of Terminal Glycosylation.Cancers (Basel). 2021 Oct 21;13(21):5273. doi: 10.3390/cancers13215273. Cancers (Basel). 2021. PMID: 34771437 Free PMC article. Review.
References
-
- American Cancer Society Cancer Facts & Figures. [(accessed on 14 January 2021)];2021 Available online: https://www.cancer.org/research/cancer-facts-statistics/all-cancer-facts....
-
- National Breast Cancer Foundation Breast Cancer Statistics. [(accessed on 13 January 2021)];2020 Available online: https://www.nationalbreastcancer.org/wp-content/uploads/2020-Breast-Canc....
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

