Intra-Colonial Viral Infections in Western Honey Bees (Apis Mellifera)
- PMID: 34070128
- PMCID: PMC8158351
- DOI: 10.3390/microorganisms9051087
Intra-Colonial Viral Infections in Western Honey Bees (Apis Mellifera)
Abstract
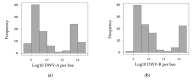
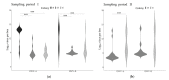
RNA viruses play a significant role in the current high losses of pollinators. Although many studies have focused on the epidemiology of western honey bee (Apis mellifera) viruses at the colony level, the dynamics of virus infection within colonies remains poorly explored. In this study, the two main variants of the ubiquitous honey bee virus DWV as well as three major honey bee viruses (SBV, ABPV and BQCV) were analyzed from Varroa-destructor-parasitized pupae. More precisely, RT-qPCR was used to quantify and compare virus genome copies across honey bee pupae at the individual and subfamily levels (i.e., patrilines, sharing the same mother queen but with different drones as fathers). Additionally, virus genome copies were compared in cells parasitized by reproducing and non-reproducing mite foundresses to assess the role of this vector. Only DWV was detected in the samples, and the two variants of this virus significantly differed when comparing the sampling period, colonies and patrilines. Moreover, DWV-A and DWV-B exhibited different infection patterns, reflecting contrasting dynamics. Altogether, these results provide new insight into honey bee diseases and stress the need for more studies about the mechanisms of intra-colonial disease variation in social insects.
Keywords: evolutionary biology; host–pathogen interactions; pollinators; population genetics; viruses.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
Grants and funding
LinkOut - more resources
Full Text Sources

