Sendai Virus-Vectored Vaccines That Express Envelope Glycoproteins of Respiratory Viruses
- PMID: 34072332
- PMCID: PMC8230104
- DOI: 10.3390/v13061023
Sendai Virus-Vectored Vaccines That Express Envelope Glycoproteins of Respiratory Viruses
Abstract
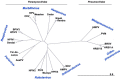
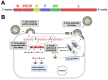
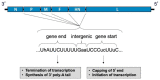
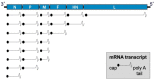
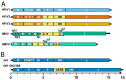
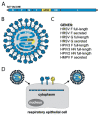
Human respiratory syncytial virus (HRSV), human metapneumovirus (HMPV), and human parainfluenza viruses (HPIVs) are leading causes of respiratory disease in young children, the elderly, and individuals of all ages with immunosuppression. Vaccination strategies against these pneumoviruses and paramyxoviruses are vast in number, yet no licensed vaccines are available. Here, we review development of Sendai virus (SeV), a versatile pediatric vaccine that can (a) serve as a Jennerian vaccine against HPIV1, (b) serve as a recombinant vaccine against HRSV, HPIV2, HPIV3, and HMPV, (c) accommodate foreign genes for viral glycoproteins in multiple intergenic positions, (d) induce durable, mucosal, B-cell, and T-cell immune responses without enhanced immunopathology, (e) protect cotton rats, African green monkeys, and chimpanzees from infection, and (f) be formulated into a vaccine cocktail. Clinical phase I safety trials of SeV have been completed in adults and 3-6-year-old children. Clinical testing of SeVRSV, an HRSV fusion (F) glycoprotein gene recombinant, has also been completed in adults. Positive results from these studies, and collaborative efforts with the National Institutes of Health and the Serum Institute of India assist advanced development of SeV-based vaccines. Prospects are now good for vaccine successes in infants and consequent protection against serious viral disease.
Keywords: HRSV; attachment protein; envelope glycoprotein; fusion glycoprotein; parainfluenza virus; paramyxovirus; pneumovirus; vaccine vector.
Conflict of interest statement
J.L.H. has received funding from NIH/NIAID research grants R01AI088729 and P01 AI054955 for SeV vaccine development. C.J.R. has received NIH/NIAID research grant R01AI083370 in part to develop the SeV vector. The work completed in the studies and the views expressed here do not necessarily represent the official views of the National Institutes of Health. Both authors are inventors of patent US2014/0186397, Modified SeV vaccine and imaging vector (3 July 2014). Additional funding to both authors was from NCI P30 CA21765 and ALSAC.
Figures
References
-
- The Global Impact of Respiratory Disease. 2nd ed. European Respiratory Society; Sheffield, UK: 2017. [(accessed on 3 March 2021)]. Forum of international respiratory societies. Available online: https://www.who.int.
-
- Johns Hopkins University and Medicine Coronavirus Resource Center. [(accessed on 6 March 2021)]; Available online: https://coronavirus.jhu.edu.
-
- Influenza 1918 Pandemic. [(accessed on 1 May 2021)]; Available online: https://www.cdc.gov.
-
- Nair H., Brooks W.A., Katz M., Roca A., Berkley J.A., Madhi S.A., Simmerman J.M., Gordon A., Sato M., Howie S., et al. Global burden of respiratory infections due to seasonal influenza in young children: A systematic review and meta-analysis. Lancet. 2011;378:1917–1930. doi: 10.1016/S0140-6736(11)61051-9. - DOI - PubMed
-
- Iuliano A.D., Roguski K.M., Chang H.H., Muscatello D.J., Palekar R., Tempia S., Cohen C., Gran J.M., Schanzer D., Cowling B.J., et al. Global Seasonal Influenza-associated Mortality Collaborator, N., Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet. 2018;391:1285–1300. doi: 10.1016/S0140-6736(17)33293-2. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

