Nitrate- and Nitrite-Sensing Histidine Kinases: Function, Structure, and Natural Diversity
- PMID: 34072989
- PMCID: PMC8199190
- DOI: 10.3390/ijms22115933
Nitrate- and Nitrite-Sensing Histidine Kinases: Function, Structure, and Natural Diversity
Abstract
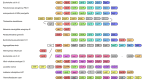
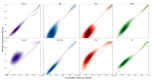
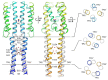
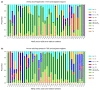
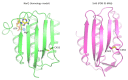
Under anaerobic conditions, bacteria may utilize nitrates and nitrites as electron acceptors. Sensitivity to nitrous compounds is achieved via several mechanisms, some of which rely on sensor histidine kinases (HKs). The best studied nitrate- and nitrite-sensing HKs (NSHKs) are NarQ and NarX from Escherichia coli. Here, we review the function of NSHKs, analyze their natural diversity, and describe the available structural information. In particular, we show that around 6000 different NSHK sequences forming several distinct clusters may now be found in genomic databases, comprising mostly the genes from Beta- and Gammaproteobacteria as well as from Bacteroidetes and Chloroflexi, including those from anaerobic ammonia oxidation (annamox) communities. We show that the architecture of NSHKs is mostly conserved, although proteins from Bacteroidetes lack the HAMP and GAF-like domains yet sometimes have PAS. We reconcile the variation of NSHK sequences with atomistic models and pinpoint the structural elements important for signal transduction from the sensor domain to the catalytic module over the transmembrane and cytoplasmic regions spanning more than 200 Å.
Keywords: allostery; cell signaling; histidine kinases; nitrate regulation; nitrate respiration; signal transduction; two-component systems.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures
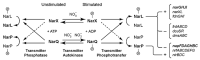








References
-
- Wassenaar T.M., Wanchai V., Alkam D., Nookaew I., Ussery D.W. Conservation of two-component signal transduction systems in E. coli, Salmonella, and Across 100,000 bacteria of various bacterial Phyla. In: Rampelotto P.H., editor. Molecular Mechanisms of Microbial Evolution. Springer International Publishing; Cham, Switzerland: 2018. pp. 153–174. Grand Challenges in Biology and, Biotechnology.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

