Innovative Approach to Fast Electron Microscopy Using the Example of a Culture of Virus-Infected Cells: An Application to SARS-CoV-2
- PMID: 34073053
- PMCID: PMC8228702
- DOI: 10.3390/microorganisms9061194
Innovative Approach to Fast Electron Microscopy Using the Example of a Culture of Virus-Infected Cells: An Application to SARS-CoV-2
Abstract
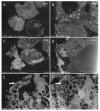
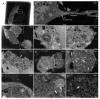
Despite the development of new diagnostic methods, co-culture, based on sample inoculation of cell monolayers coupled with electron microscopy (EM) observation, remains the gold standard in virology. Indeed, co-culture allows for the study of cell morphology (infected and not infected), the ultrastructure of the inoculated virus, and the different steps of the virus infectious cycle. Most EM methods for studying virus cycles are applied after infected cells are produced in large quantities and detached to obtain a pellet. Here, cell culture was performed in sterilized, collagen-coated single-break strip wells. After one day in culture, cells were infected with SARS-CoV-2. Wells of interest were fixed at different time points, from 2 to 36 h post-infection. Microwave-assisted resin embedding was accomplished directly in the wells in 4 h. Finally, ultra-thin sections were cut directly through the infected-cell monolayers. Our methodology requires, in total, less than four days for preparing and observing cells. Furthermore, by observing undetached infected cell monolayers, we were able to observe new ultrastructural findings, such as cell-cell interactions and baso-apical cellular organization related to the virus infectious cycle. Our innovative methodology thus not only saves time for preparation but also adds precision and new knowledge about viral infection, as shown here for SARS-CoV-2.
Keywords: SARS-CoV-2; electron microscopy; embedding method; microwaves; single-break strip.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Goldsmith C.S., Ksiazek T.G., Rollin P.E., Comer J.A., Nicholson W.L., Peret T.C.T., Erdman D.D., Bellini W.J., Harcourt B.H., Rota P.A., et al. Cell Culture and Electron Microscopy for Identifying Viruses in Diseases of Unknown Cause. Emerg. Infect. Dis. 2013;19:886–891. doi: 10.3201/eid1906.130173. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

