Advances and Perspectives in Tissue Culture and Genetic Engineering of Cannabis
- PMID: 34073522
- PMCID: PMC8197860
- DOI: 10.3390/ijms22115671
Advances and Perspectives in Tissue Culture and Genetic Engineering of Cannabis
Abstract
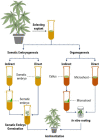
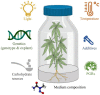
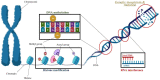
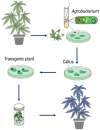
For a long time, Cannabis sativa has been used for therapeutic and industrial purposes. Due to its increasing demand in medicine, recreation, and industry, there is a dire need to apply new biotechnological tools to introduce new genotypes with desirable traits and enhanced secondary metabolite production. Micropropagation, conservation, cell suspension culture, hairy root culture, polyploidy manipulation, and Agrobacterium-mediated gene transformation have been studied and used in cannabis. However, some obstacles such as the low rate of transgenic plant regeneration and low efficiency of secondary metabolite production in hairy root culture and cell suspension culture have restricted the application of these approaches in cannabis. In the current review, in vitro culture and genetic engineering methods in cannabis along with other promising techniques such as morphogenic genes, new computational approaches, clustered regularly interspaced short palindromic repeats (CRISPR), CRISPR/Cas9-equipped Agrobacterium-mediated genome editing, and hairy root culture, that can help improve gene transformation and plant regeneration, as well as enhance secondary metabolite production, have been highlighted and discussed.
Keywords: gene transformation; genome editing; haploid production; hemp; in vitro culture; marijuana; morphogenic genes; organogenesis; polyploidy; somatic embryogenesis.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures


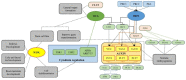
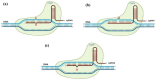
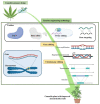
References
-
- Hesami M., Pepe M., Alizadeh M., Rakei A., Baiton A., Jones A.M.P. Recent advances in cannabis biotechnology. Ind. Crop. Prod. 2020;158:113026. doi: 10.1016/j.indcrop.2020.113026. - DOI
-
- Schultz C.J., Lim W.L., Khor S.F., Neumann K.A., Schulz J.M., Ansari O., Skewes M.A., Burton R.A. Consumer and health-related traits of seed from selected commercial and breeding lines of industrial hemp, Cannabis sativa L. J. Agric. Food Res. 2020;2:100025. doi: 10.1016/j.jafr.2020.100025. - DOI
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

