p62 overexpression induces TDP-43 cytoplasmic mislocalisation, aggregation and cleavage and neuronal death
- PMID: 34075102
- PMCID: PMC8169680
- DOI: 10.1038/s41598-021-90822-2
p62 overexpression induces TDP-43 cytoplasmic mislocalisation, aggregation and cleavage and neuronal death
Abstract
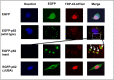
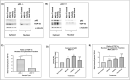
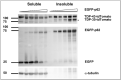
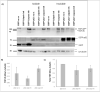
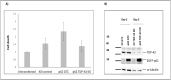
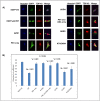
Amyotrophic lateral sclerosis (ALS) and frontotemporal lobar degeneration (FTLD) that exist on a spectrum of neurodegenerative disease. A hallmark of pathology is cytoplasmic TDP-43 aggregates within neurons, observed in 97% of ALS cases and ~ 50% of FTLD cases. This mislocalisation from the nucleus into the cytoplasm and TDP-43 cleavage are associated with pathology, however, the drivers of these changes are unknown. p62 is invariably also present within these aggregates. We show that p62 overexpression causes TDP-43 mislocalisation into cytoplasmic aggregates, and aberrant TDP-43 cleavage that was dependent on both the PB1 and ubiquitin-associated (UBA) domains of p62. We further show that p62 overexpression induces neuron death. We found that stressors (proteasome inhibition and arsenic) increased p62 expression and that this shifted the nuclear:cytoplasmic TDP-43 ratio. Overall, our study suggests that environmental factors that increase p62 may thereby contribute to TDP-43 pathology in ALS and FTLD.
Conflict of interest statement
The authors declare no competing interests.
Figures






Similar articles
-
p62 positive, TDP-43 negative, neuronal cytoplasmic and intranuclear inclusions in the cerebellum and hippocampus define the pathology of C9orf72-linked FTLD and MND/ALS.Acta Neuropathol. 2011 Dec;122(6):691-702. doi: 10.1007/s00401-011-0911-2. Epub 2011 Nov 19. Acta Neuropathol. 2011. PMID: 22101323
-
TDP-43 transgenic mice develop spastic paralysis and neuronal inclusions characteristic of ALS and frontotemporal lobar degeneration.Proc Natl Acad Sci U S A. 2010 Feb 23;107(8):3858-63. doi: 10.1073/pnas.0912417107. Epub 2010 Feb 3. Proc Natl Acad Sci U S A. 2010. PMID: 20133711 Free PMC article.
-
TDP-43 is consistently co-localized with ubiquitinated inclusions in sporadic and Guam amyotrophic lateral sclerosis but not in familial amyotrophic lateral sclerosis with and without SOD1 mutations.Neuropathology. 2009 Dec;29(6):672-83. doi: 10.1111/j.1440-1789.2009.01029.x. Epub 2009 Jun 3. Neuropathology. 2009. PMID: 19496940
-
Pathway from TDP-43-Related Pathology to Neuronal Dysfunction in Amyotrophic Lateral Sclerosis and Frontotemporal Lobar Degeneration.Int J Mol Sci. 2021 Apr 8;22(8):3843. doi: 10.3390/ijms22083843. Int J Mol Sci. 2021. PMID: 33917673 Free PMC article. Review.
-
TDP-43 proteinopathy in frontotemporal lobar degeneration and amyotrophic lateral sclerosis: From pathomechanisms to therapeutic strategies.Ageing Res Rev. 2024 Sep;100:102441. doi: 10.1016/j.arr.2024.102441. Epub 2024 Jul 27. Ageing Res Rev. 2024. PMID: 39069095 Review.
Cited by
-
Effects of intracellular calcium accumulation on proteins encoded by the major genes underlying amyotrophic lateral sclerosis.Sci Rep. 2022 Jan 10;12(1):395. doi: 10.1038/s41598-021-04267-8. Sci Rep. 2022. PMID: 35013445 Free PMC article.
-
Survival in sporadic ALS is associated with lower p62 burden in the spinal cord.J Neuropathol Exp Neurol. 2023 Aug 21;82(9):769-773. doi: 10.1093/jnen/nlad051. J Neuropathol Exp Neurol. 2023. PMID: 37414530 Free PMC article.
-
Proximity labeling reveals dynamic changes in the SQSTM1 protein network.bioRxiv [Preprint]. 2024 Jun 27:2023.12.12.571324. doi: 10.1101/2023.12.12.571324. bioRxiv. 2024. Update in: J Biol Chem. 2024 Sep;300(9):107621. doi: 10.1016/j.jbc.2024.107621. PMID: 38168279 Free PMC article. Updated. Preprint.
-
Loss of TDP-43 oligomerization or RNA binding elicits distinct aggregation patterns.EMBO J. 2023 Sep 4;42(17):e111719. doi: 10.15252/embj.2022111719. Epub 2023 Jul 11. EMBO J. 2023. PMID: 37431963 Free PMC article.
-
The Molecular Intersection of NEK1, C21ORF2, Cyclin F, and VCP in ALS Pathogenesis.Genes (Basel). 2025 Mar 30;16(4):407. doi: 10.3390/genes16040407. Genes (Basel). 2025. PMID: 40282367 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

