Dot/Icm-Dependent Restriction of Legionella pneumophila within Neutrophils
- PMID: 34076467
- PMCID: PMC8262857
- DOI: 10.1128/mBio.01008-21
Dot/Icm-Dependent Restriction of Legionella pneumophila within Neutrophils
Abstract
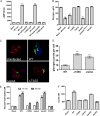
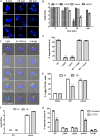
The Dot/Icm type IV secretion system (T4SS) of Legionella pneumophila is essential for lysosomal evasion and permissiveness of macrophages for intracellular proliferation of the pathogen. In contrast, we show that polymorphonuclear cells (PMNs) respond to a functional Dot/Icm system through rapid restriction of L. pneumophila. Specifically, we show that the L. pneumophila T4SS-injected amylase (LamA) effector catalyzes rapid glycogen degradation in the PMNs cytosol, leading to cytosolic hyperglucose. Neutrophils respond through immunometabolic reprogramming that includes upregulated aerobic glycolysis. The PMNs become activated with spatial generation of intracellular reactive oxygen species within the Legionella-containing phagosome (LCP) and fusion of specific and azurophilic granules to the LCP, leading to rapid restriction of L. pneumophila. We conclude that in contrast to macrophages, PMNs respond to a functional Dot/Icm system, and specifically to the effect of the injected amylase effector, through rapid engagement of major microbicidal processes and rapid restriction of the pathogen. IMPORTANCE Legionella pneumophila is commonly found in aquatic environments and resides within a wide variety of amoebal hosts. Upon aerosol transmission to humans, L. pneumophila invades and replicates with alveolar macrophages, causing pneumonia designated Legionnaires' disease. In addition to alveolar macrophages, neutrophils infiltrate into the lungs of infected patients. Unlike alveolar macrophages, neutrophils restrict and kill L. pneumophila, but the mechanisms were previously unclear. Here, we show that the pathogen secretes an amylase (LamA) enzyme that rapidly breakdowns glycogen stores within neutrophils, and this triggers increased glycolysis. Subsequently, the two major killing mechanisms of neutrophils, granule fusion and production of reactive oxygen species, are activated, resulting in rapid killing of L. pneumophila.
Keywords: granules; phagosome; polymorphonuclear leukocytes; reactive oxygen species.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

