Mobile Oxazolidinone Resistance Genes in Gram-Positive and Gram-Negative Bacteria
- PMID: 34076490
- PMCID: PMC8262807
- DOI: 10.1128/CMR.00188-20
Mobile Oxazolidinone Resistance Genes in Gram-Positive and Gram-Negative Bacteria
Abstract
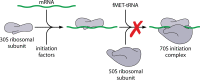
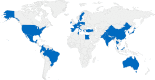
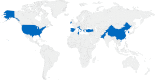
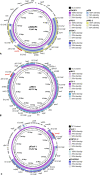
Seven mobile oxazolidinone resistance genes, including cfr, cfr(B), cfr(C), cfr(D), cfr(E), optrA, and poxtA, have been identified to date. The cfr genes code for 23S rRNA methylases, which confer a multiresistance phenotype that includes resistance to phenicols, lincosamides, oxazolidinones, pleuromutilins, and streptogramin A compounds. The optrA and poxtA genes code for ABC-F proteins that protect the bacterial ribosomes from the inhibitory effects of oxazolidinones. The optrA gene confers resistance to oxazolidinones and phenicols, while the poxtA gene confers elevated MICs or resistance to oxazolidinones, phenicols, and tetracycline. These oxazolidinone resistance genes are most frequently found on plasmids, but they are also located on transposons, integrative and conjugative elements (ICEs), genomic islands, and prophages. In these mobile genetic elements (MGEs), insertion sequences (IS) most often flanked the cfr, optrA, and poxtA genes and were able to generate translocatable units (TUs) that comprise the oxazolidinone resistance genes and occasionally also other genes. MGEs and TUs play an important role in the dissemination of oxazolidinone resistance genes across strain, species, and genus boundaries. Most frequently, these MGEs also harbor genes that mediate resistance not only to antimicrobial agents of other classes, but also to metals and biocides. Direct selection pressure by the use of antimicrobial agents to which the oxazolidinone resistance genes confer resistance, but also indirect selection pressure by the use of antimicrobial agents, metals, or biocides (the respective resistance genes against which are colocated on cfr-, optrA-, or poxtA-carrying MGEs) may play a role in the coselection and persistence of oxazolidinone resistance genes.
Keywords: cfr; genomic island; horizontal transfer; integrative and conjugative element; mobile genetic element; optrA; oxazolidinones; plasmid; poxtA; prophage; transposon.
Figures







References
-
- Slee AM, Wuonola MA, McRipley RJ, Zajac I, Zawada MJ, Bartholomew PT, Gregory WA, Forbes M. 1987. Oxazolidinones, a new class of synthetic antibacterial agents: in vitro and in vivo activities of DuP 105 and DuP 721. Antimicrob Agents Chemother 31:1791–1797. 10.1128/aac.31.11.1791. - DOI - PMC - PubMed
-
- Brickner SJ, Hutchinson DK, Barbachyn MR, Manninen PR, Ulanowicz DA, Garmon SA, Grega KC, Hendges SK, Toops DS, Ford CW, Zurenko GE. 1996. Synthesis and antibacterial activity of U-100592 and U-100766, two oxazolidinone antibacterial agents for the potential treatment of multidrug-resistant Gram-positive bacterial infections. J Med Chem 39:673–679. 10.1021/jm9509556. - DOI - PubMed
-
- Tsuji BT, Kaatz GW, Ryback MJ. 1999. –2021. Linezolid and other oxazolidinones. In Antimicrobial therapy and vaccines volume II: antimicrobial agents. Antimicrobe, Pittsburgh, PA. http://www.antimicrobe.org/d13.asp. Accessed 12 Jan 2021.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

