Modelling covalent linkages in CCP4
- PMID: 34076587
- PMCID: PMC8171069
- DOI: 10.1107/S2059798321001753
Modelling covalent linkages in CCP4
Abstract
In this contribution, the current protocols for modelling covalent linkages within the CCP4 suite are considered. The mechanism used for modelling covalent linkages is reviewed: the use of dictionaries for describing changes to stereochemistry as a result of the covalent linkage and the application of link-annotation records to structural models to ensure the correct treatment of individual instances of covalent linkages. Previously, linkage descriptions were lacking in quality compared with those of contemporary component dictionaries. Consequently, AceDRG has been adapted for the generation of link dictionaries of the same quality as for individual components. The approach adopted by AceDRG for the generation of link dictionaries is outlined, which includes associated modifications to the linked components. A number of tools to facilitate the practical modelling of covalent linkages available within the CCP4 suite are described, including a new restraint-dictionary accumulator, the Make Covalent Link tool and AceDRG interface in Coot, the 3D graphical editor JLigand and the mechanisms for dealing with covalent linkages in the CCP4i2 and CCP4 Cloud environments. These integrated solutions streamline and ease the covalent-linkage modelling workflow, seamlessly transferring relevant information between programs. Current recommended practice is elucidated by means of instructive practical examples. By summarizing the different approaches to modelling linkages that are available within the CCP4 suite, limitations and potential pitfalls that may be encountered are highlighted in order to raise awareness, with the intention of improving the quality of future modelled covalent linkages in macromolecular complexes.
Keywords: AceDRG; CCP4; CCP4 Monomer Library; covalent linkages; link dictionary; link records; link-restraint dictionary; mmCIF; monomer library; restraints.
open access.
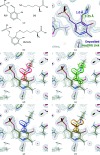
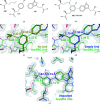
Figures
Similar articles
-
The missing link: covalent linkages in structural models.Acta Crystallogr D Struct Biol. 2021 Jun 1;77(Pt 6):727-745. doi: 10.1107/S2059798321003934. Epub 2021 May 19. Acta Crystallogr D Struct Biol. 2021. PMID: 34076588 Free PMC article.
-
CCP4i2: the new graphical user interface to the CCP4 program suite.Acta Crystallogr D Struct Biol. 2018 Feb 1;74(Pt 2):68-84. doi: 10.1107/S2059798317016035. Epub 2018 Feb 1. Acta Crystallogr D Struct Biol. 2018. PMID: 29533233 Free PMC article.
-
Ligand fitting with CCP4.Acta Crystallogr D Struct Biol. 2017 Feb 1;73(Pt 2):158-170. doi: 10.1107/S2059798316020143. Epub 2017 Feb 1. Acta Crystallogr D Struct Biol. 2017. PMID: 28177312 Free PMC article.
-
Ongoing developments in CCP4 for high-throughput structure determination.Acta Crystallogr D Biol Crystallogr. 2002 Nov;58(Pt 11):1929-36. doi: 10.1107/s0907444902016116. Epub 2002 Oct 21. Acta Crystallogr D Biol Crystallogr. 2002. PMID: 12393924 Review.
-
The Protein Data Bank Archive.Methods Mol Biol. 2021;2305:3-21. doi: 10.1007/978-1-0716-1406-8_1. Methods Mol Biol. 2021. PMID: 33950382 Review.
Cited by
-
The missing link: covalent linkages in structural models.Acta Crystallogr D Struct Biol. 2021 Jun 1;77(Pt 6):727-745. doi: 10.1107/S2059798321003934. Epub 2021 May 19. Acta Crystallogr D Struct Biol. 2021. PMID: 34076588 Free PMC article.
-
The CCP4 suite: integrative software for macromolecular crystallography.Acta Crystallogr D Struct Biol. 2023 Jun 1;79(Pt 6):449-461. doi: 10.1107/S2059798323003595. Epub 2023 May 30. Acta Crystallogr D Struct Biol. 2023. PMID: 37259835 Free PMC article.
-
Neutron crystallographic refinement with REFMAC5 from the CCP4 suite.Acta Crystallogr D Struct Biol. 2023 Dec 1;79(Pt 12):1056-1070. doi: 10.1107/S2059798323008793. Epub 2023 Nov 3. Acta Crystallogr D Struct Biol. 2023. PMID: 37921806 Free PMC article.
-
Crystallographic fragment-binding studies of the Mycobacterium tuberculosis trifunctional enzyme suggest binding pockets for the tails of the acyl-CoA substrates at its active sites and a potential substrate-channeling path between them.Acta Crystallogr D Struct Biol. 2024 Aug 1;80(Pt 8):605-619. doi: 10.1107/S2059798324006557. Epub 2024 Jul 16. Acta Crystallogr D Struct Biol. 2024. PMID: 39012716 Free PMC article.
-
Specific tRNAs promote mRNA decay by recruiting the CCR4-NOT complex to translating ribosomes.Science. 2024 Nov 22;386(6724):eadq8587. doi: 10.1126/science.adq8587. Epub 2024 Nov 22. Science. 2024. PMID: 39571015 Free PMC article.
References
-
- Adams, P. D., Afonine, P. V., Baskaran, K., Berman, H. M., Berrisford, J., Bricogne, G., Brown, D. G., Burley, S. K., Chen, M., Feng, Z., Flensburg, C., Gutmanas, A., Hoch, J. C., Ikegawa, Y., Kengaku, Y., Krissinel, E., Kurisu, G., Liang, Y., Liebschner, D., Mak, L., Markley, J. L., Moriarty, N. W., Murshudov, G. N., Noble, M., Peisach, E., Persikova, I., Poon, B. K., Sobolev, O. V., Ulrich, E. L., Velankar, S., Vonrhein, C., Westbrook, J., Wojdyr, M., Yokochi, M. & Young, J. Y. (2019). Acta Cryst. D75, 451–454. - PMC - PubMed
-
- Bertrand, T., Eady, N. A., Jones, J. N., Jesmin, Nagy, J. M., Jamart-Grégoire, B., Raven, E. L. & Brown, K. A. (2004). J. Biol. Chem. 279, 38991–38999. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

