Cannabinoid Signaling in Auditory Function and Development
- PMID: 34079440
- PMCID: PMC8165240
- DOI: 10.3389/fnmol.2021.678510
Cannabinoid Signaling in Auditory Function and Development
Abstract
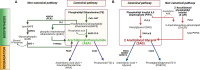
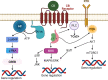
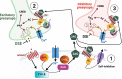
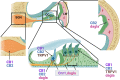
Plants of the genus Cannabis have been used by humans for millennia for a variety of purposes. Perhaps most notable is the use of certain Cannabis strains for their psychoactive effects. More recently, several biologically active molecules within the plants of these Cannabis strains, called phytocannabinoids or simply cannabinoids, have been identified. Furthermore, within human cells, endogenous cannabinoids, or endocannabinoids, as well as the receptors and secondary messengers that give rise to their neuromodulatory effects, have also been characterized. This endocannabinoid system (ECS) is composed of two primary ligands-anandamide and 2-arachidonyl glycerol; two primary receptors-cannabinoid receptors 1 and 2; and several enzymes involved in biosynthesis and degradation of endocannabinoid ligands including diacylglycerol lipase (DAGL) and monoacylglycerol lipase (MAGL). Here we briefly summarize cannabinoid signaling and review what has been discerned to date with regard to cannabinoid signaling in the auditory system and its roles in normal physiological function as well as pathological conditions. While much has been uncovered regarding cannabinoid signaling in the central nervous system, less attention has been paid to the auditory system specifically. Still, evidence is emerging to suggest that cannabinoid signaling is critical for the development, maturation, function, and survival of cochlear hair cells (HCs) and spiral ganglion neurons (SGNs). Furthermore, cannabinoid signaling can have profound effects on synaptic connectivity in CNS structures related to auditory processing. While clinical cases demonstrate that endogenous and exogenous cannabinoids impact auditory function, this review highlights several areas, such as SGN development, where more research is warranted.
Keywords: cannabinoid; cochlea; hair cell; hearing; hearing—drug effects; otoprotection; spiral ganglion.
Copyright © 2021 Ghosh, Stansak and Walters.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
The Endocannabinoid System Modulating Levels of Consciousness, Emotions and Likely Dream Contents.CNS Neurol Disord Drug Targets. 2017;16(4):370-379. doi: 10.2174/1871527316666170223161908. CNS Neurol Disord Drug Targets. 2017. PMID: 28240187 Review.
-
Review of the Endocannabinoid System.Biol Psychiatry Cogn Neurosci Neuroimaging. 2021 Jun;6(6):607-615. doi: 10.1016/j.bpsc.2020.07.016. Epub 2020 Aug 1. Biol Psychiatry Cogn Neurosci Neuroimaging. 2021. PMID: 32980261 Free PMC article. Review.
-
An Introduction to the Endogenous Cannabinoid System.Biol Psychiatry. 2016 Apr 1;79(7):516-25. doi: 10.1016/j.biopsych.2015.07.028. Epub 2015 Oct 30. Biol Psychiatry. 2016. PMID: 26698193 Free PMC article. Review.
-
Cannabinoid receptor signaling in progenitor/stem cell proliferation and differentiation.Prog Lipid Res. 2013 Oct;52(4):633-50. doi: 10.1016/j.plipres.2013.05.004. Epub 2013 Sep 25. Prog Lipid Res. 2013. PMID: 24076098 Review.
-
Phytocannabinoids for Cancer Therapeutics: Recent Updates and Future Prospects.Curr Med Chem. 2015;22(30):3472-501. doi: 10.2174/0929867322666150716115057. Curr Med Chem. 2015. PMID: 26179998 Review.
Cited by
-
Plasma metabolomics analyses highlight the multifaceted effects of noise exposure and the diagnostic power of dysregulated metabolites for noise-induced hearing loss in steel workers.Front Mol Biosci. 2022 Aug 19;9:907832. doi: 10.3389/fmolb.2022.907832. eCollection 2022. Front Mol Biosci. 2022. PMID: 36060246 Free PMC article.
-
Cannabidiol regulates apoptosis and autophagy in inflammation and cancer: A review.Front Pharmacol. 2023 Jan 23;14:1094020. doi: 10.3389/fphar.2023.1094020. eCollection 2023. Front Pharmacol. 2023. PMID: 36755953 Free PMC article. Review.
-
Potential uses of auditory nerve stimulation to modulate immune responses in the inner ear and auditory brainstem.Front Integr Neurosci. 2023 Dec 14;17:1294525. doi: 10.3389/fnint.2023.1294525. eCollection 2023. Front Integr Neurosci. 2023. PMID: 38162822 Free PMC article.
-
A Systematic Review on Metabolomics Analysis in Hearing Impairment: Is It a Possible Tool in Understanding Auditory Pathologies?Int J Mol Sci. 2023 Oct 14;24(20):15188. doi: 10.3390/ijms242015188. Int J Mol Sci. 2023. PMID: 37894867 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources

