Tight nanoscale clustering of Fcγ receptors using DNA origami promotes phagocytosis
- PMID: 34080973
- PMCID: PMC8175083
- DOI: 10.7554/eLife.68311
Tight nanoscale clustering of Fcγ receptors using DNA origami promotes phagocytosis
Abstract
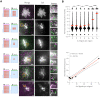
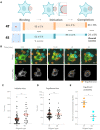
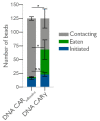
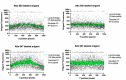
Macrophages destroy pathogens and diseased cells through Fcγ receptor (FcγR)-driven phagocytosis of antibody-opsonized targets. Phagocytosis requires activation of multiple FcγRs, but the mechanism controlling the threshold for response is unclear. We developed a DNA origami-based engulfment system that allows precise nanoscale control of the number and spacing of ligands. When the number of ligands remains constant, reducing ligand spacing from 17.5 nm to 7 nm potently enhances engulfment, primarily by increasing efficiency of the engulfment-initiation process. Tighter ligand clustering increases receptor phosphorylation, as well as proximal downstream signals. Increasing the number of signaling domains recruited to a single ligand-receptor complex was not sufficient to recapitulate this effect, indicating that clustering of multiple receptors is required. Our results suggest that macrophages use information about local ligand densities to make critical engulfment decisions, which has implications for the mechanism of antibody-mediated phagocytosis and the design of immunotherapies.
Keywords: Antibody; DNA origami; Fc Receptor; Phagocytosis; cell biology; human; immunology; immunotherapy; inflammation; mouse; synthetic biology.
Plain language summary
The word ‘phagocytosis’ means cellular eating. It is the process by which cells extend their membranes around foreign particles and engulf them. Macrophages, a type of immune cell found in every tissue of the body, perform phagocytosis to eat pathogens and diseased cells. To avoid eating healthy cells, macrophages focus on targets marked by proteins called antibodies. They look for cells coated with high levels of a type of antibody called immunoglobulin G, or IgG for short, but only eat cells coated with enough IgG, raising the question, can macrophages count? Macrophages recognize IgG antibodies using cell surface receptors called Fc-gamma Receptors. When these receptors bind to IgG, they cluster together. Researchers do not yet know how the number of IgG antibodies per cluster, or the spacing between them, affects phagocytosis. To find this out, researchers need to be able to manipulate the clustering experimentally. One way to do this is using a technique called DNA origami. This technique creates nanoscale patterns of DNA strands on a target surface. If the part of a receptor that interacts with its target is then replaced with a complementary DNA strand to the strands on the target surface, the receptor will bind the surface following the nanoscale pattern. This allows researchers to generate synthetic targets with specific patterns of receptor-target interaction. Kern et al. replaced the part of the macrophage Fc-gamma Receptor that interacts with IgG with a strand of DNA. They then used DNA origami to arrange complementary DNA strands on pegboards and attached these pegboards to silica beads. The different arrangements of DNA on these pegboards mimicked the types of antibody clusters macrophages might encounter on the surfaces of the cells and particles they have to engulf in the body. Kern et al. found that tight clusters of the DNA targets on the pegboards made the macrophages most likely to begin phagocytosis, particularly clusters of eight or more DNA strands spaced less than seven nanometers apart. Macrophages encountering these tight clusters showed an increase in Fc-gamma receptor activation, which is crucial for macrophage attack. Whether or not macrophages can count, they can at least sense the level of clustering of IgG antibodies to determine if a target should be engulfed. Doctors use antibody therapies that rely on Fc-gamma receptor engagement to treat cancer, autoimmune and neurodegenerative diseases. Understanding how clustering affects phagocytosis could aid in the design of new antibody treatments. It could also help improve the design of synthetic receptors to create designer immune cells that can attack specific targets. The next step will be to recreate the results from the synthetic system used by Kern et al. with natural receptors and antibodies.
© 2021, Kern et al.
Conflict of interest statement
NK, RD, SD, RV, MM No competing interests declared
Figures