Historical genomics reveals the evolutionary mechanisms behind multiple outbreaks of the host-specific coffee wilt pathogen Fusarium xylarioides
- PMID: 34082717
- PMCID: PMC8176585
- DOI: 10.1186/s12864-021-07700-4
Historical genomics reveals the evolutionary mechanisms behind multiple outbreaks of the host-specific coffee wilt pathogen Fusarium xylarioides
Erratum in
-
Correction to: Historical genomics reveals the evolutionary mechanisms behind multiple outbreaks of the host-specific coffee wilt pathogen Fusarium xylarioides.BMC Genomics. 2021 Jul 7;22(1):514. doi: 10.1186/s12864-021-07831-8. BMC Genomics. 2021. PMID: 34233621 Free PMC article. No abstract available.
Abstract
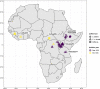
Background: Nearly 50% of crop yields are lost to pests and disease, with plants and pathogens locked in an amplified co-evolutionary process of disease outbreaks. Coffee wilt disease, caused by Fusarium xylarioides, decimated coffee production in west and central Africa following its initial outbreak in the 1920s. After successful management, it later re-emerged and by the 2000s comprised two separate epidemics on arabica coffee in Ethiopia and robusta coffee in east and central Africa.
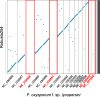
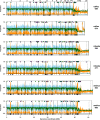
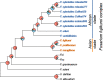
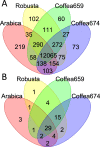
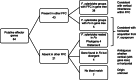
Results: Here, we use genome sequencing of six historical culture collection strains spanning 52 years to identify the evolutionary processes behind these repeated outbreaks. Phylogenomic reconstruction using 13,782 single copy orthologs shows that the robusta population arose from the initial outbreak, whilst the arabica population is a divergent sister clade to the other strains. A screen for putative effector genes involved in pathogenesis shows that the populations have diverged in gene content and sequence mainly by vertical processes within lineages. However, 15 putative effector genes show evidence of horizontal acquisition, with close homology to genes from F. oxysporum. Most occupy small regions of homology within wider scaffolds, whereas a cluster of four genes occupy a 20Kb scaffold with strong homology to a region on a mobile pathogenicity chromosome in F. oxysporum that houses known effector genes. Lacking a match to the whole mobile chromosome, we nonetheless found close associations with DNA transposons, especially the miniature impala type previously proposed to facilitate horizontal transfer of pathogenicity genes in F. oxysporum. These findings support a working hypothesis that the arabica and robusta populations partly acquired distinct effector genes via transposition-mediated horizontal transfer from F. oxysporum, which shares coffee as a host and lives on other plants intercropped with coffee.
Conclusion: Our results show how historical genomics can help reveal mechanisms that allow fungal pathogens to keep pace with our efforts to resist them. Our list of putative effector genes identifies possible future targets for fungal control. In turn, knowledge of horizontal transfer mechanisms and putative donor taxa might help to design future intercropping strategies that minimize the risk of transfer of effector genes between closely-related Fusarium taxa.
Keywords: Comparative genomics; Effector; Fungi; Fusarium oxysporum; Host adaptation; Proteome.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
-
- Strange RN, Scott PR. Plant disease: a threat to global food security. Annu Rev Phytopathol. 2005;43:83–116. - PubMed
-
- Jones JDG, Dangl JL. The plant immune system. Nature. 2006;444:323–9. - PubMed
-
- Möller M, Stukenbrock EH. Evolution and genome architecture in fungal plant pathogens. Nat Rev Microbiol. 2017;15(12):756–71. - PubMed
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

