Nitrate Removal by a Novel Lithoautotrophic Nitrate-Reducing, Iron(II)-Oxidizing Culture Enriched from a Pyrite-Rich Limestone Aquifer
- PMID: 34085863
- PMCID: PMC8373248
- DOI: 10.1128/AEM.00460-21
Nitrate Removal by a Novel Lithoautotrophic Nitrate-Reducing, Iron(II)-Oxidizing Culture Enriched from a Pyrite-Rich Limestone Aquifer
Abstract
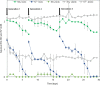
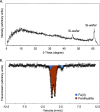
Nitrate removal in oligotrophic environments is often limited by the availability of suitable organic electron donors. Chemolithoautotrophic bacteria may play a key role in denitrification in aquifers depleted in organic carbon. Under anoxic and circumneutral pH conditions, iron(II) was hypothesized to serve as an electron donor for microbially mediated nitrate reduction by Fe(II)-oxidizing (NRFeOx) microorganisms. However, lithoautotrophic NRFeOx cultures have never been enriched from any aquifer, and as such, there are no model cultures available to study the physiology and geochemistry of this potentially environmentally relevant process. Using iron(II) as an electron donor, we enriched a lithoautotrophic NRFeOx culture from nitrate-containing groundwater of a pyrite-rich limestone aquifer. In the enriched NRFeOx culture that does not require additional organic cosubstrates for growth, within 7 to 11 days, 0.3 to 0.5 mM nitrate was reduced and 1.3 to 2 mM iron(II) was oxidized, leading to a stoichiometric NO3-/Fe(II) ratio of 0.2, with N2 and N2O identified as the main nitrate reduction products. Short-range ordered Fe(III) (oxyhydr)oxides were the product of iron(II) oxidation. Microorganisms were observed to be closely associated with formed minerals, but only few cells were encrusted, suggesting that most of the bacteria were able to avoid mineral precipitation at their surface. Analysis of the microbial community by long-read 16S rRNA gene sequencing revealed that the culture is dominated by members of the Gallionellaceae family that are known as autotrophic, neutrophilic, and microaerophilic iron(II) oxidizers. In summary, our study suggests that NRFeOx mediated by lithoautotrophic bacteria can lead to nitrate removal in anthropogenically affected aquifers. IMPORTANCE Removal of nitrate by microbial denitrification in groundwater is often limited by low concentrations of organic carbon. In these carbon-poor ecosystems, nitrate-reducing bacteria that can use inorganic compounds such as Fe(II) (NRFeOx) as electron donors could play a major role in nitrate removal. However, no lithoautotrophic NRFeOx culture has been successfully isolated or enriched from this type of environment, and as such, there are no model cultures available to study the rate-limiting factors of this potentially important process. Here, we present the physiology and microbial community composition of a novel lithoautotrophic NRFeOx culture enriched from a fractured aquifer in southern Germany. The culture is dominated by a putative Fe(II) oxidizer affiliated with the Gallionellaceae family and performs nitrate reduction coupled to Fe(II) oxidation leading to N2O and N2 formation without the addition of organic substrates. Our analyses demonstrate that lithoautotrophic NRFeOx can potentially lead to nitrate removal in nitrate-contaminated aquifers.
Keywords: NRFeOx; aquifer; geomicrobiology; groundwater; iron metabolism; iron oxidizers; nitrate; pyrite.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

