SepF is the FtsZ anchor in archaea, with features of an ancestral cell division system
- PMID: 34088904
- PMCID: PMC8178401
- DOI: 10.1038/s41467-021-23099-8
SepF is the FtsZ anchor in archaea, with features of an ancestral cell division system
Abstract
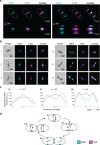
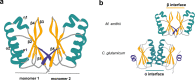
Most archaea divide by binary fission using an FtsZ-based system similar to that of bacteria, but they lack many of the divisome components described in model bacterial organisms. Notably, among the multiple factors that tether FtsZ to the membrane during bacterial cell constriction, archaea only possess SepF-like homologs. Here, we combine structural, cellular, and evolutionary analyses to demonstrate that SepF is the FtsZ anchor in the human-associated archaeon Methanobrevibacter smithii. 3D super-resolution microscopy and quantitative analysis of immunolabeled cells show that SepF transiently co-localizes with FtsZ at the septum and possibly primes the future division plane. M. smithii SepF binds to membranes and to FtsZ, inducing filament bundling. High-resolution crystal structures of archaeal SepF alone and in complex with the FtsZ C-terminal domain (FtsZCTD) reveal that SepF forms a dimer with a homodimerization interface driving a binding mode that is different from that previously reported in bacteria. Phylogenetic analyses of SepF and FtsZ from bacteria and archaea indicate that the two proteins may date back to the Last Universal Common Ancestor (LUCA), and we speculate that the archaeal mode of SepF/FtsZ interaction might reflect an ancestral feature. Our results provide insights into the mechanisms of archaeal cell division and pave the way for a better understanding of the processes underlying the divide between the two prokaryotic domains.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Structural and genetic analyses reveal the protein SepF as a new membrane anchor for the Z ring.Proc Natl Acad Sci U S A. 2013 Nov 26;110(48):E4601-10. doi: 10.1073/pnas.1313978110. Epub 2013 Nov 11. Proc Natl Acad Sci U S A. 2013. PMID: 24218584 Free PMC article.
-
The archaeal protein SepF is essential for cell division in Haloferax volcanii.Nat Commun. 2021 Jun 8;12(1):3469. doi: 10.1038/s41467-021-23686-9. Nat Commun. 2021. PMID: 34103513 Free PMC article.
-
Bacillus subtilis SepF binds to the C-terminus of FtsZ.PLoS One. 2012;7(8):e43293. doi: 10.1371/journal.pone.0043293. Epub 2012 Aug 13. PLoS One. 2012. PMID: 22912848 Free PMC article.
-
Spotlight on FtsZ-based cell division in Archaea.Trends Microbiol. 2022 Jul;30(7):665-678. doi: 10.1016/j.tim.2022.01.005. Epub 2022 Mar 1. Trends Microbiol. 2022. PMID: 35246355 Review.
-
Evolution of diverse cell division and vesicle formation systems in Archaea.Nat Rev Microbiol. 2010 Oct;8(10):731-41. doi: 10.1038/nrmicro2406. Epub 2010 Sep 6. Nat Rev Microbiol. 2010. PMID: 20818414 Free PMC article. Review.
Cited by
-
Diversity, Origin and Evolution of the ESCRT Systems.bioRxiv [Preprint]. 2024 Feb 7:2024.02.06.579148. doi: 10.1101/2024.02.06.579148. bioRxiv. 2024. Update in: mBio. 2024 Mar 13;15(3):e0033524. doi: 10.1128/mbio.00335-24. PMID: 38903064 Free PMC article. Updated. Preprint.
-
Tracing back variations in archaeal ESCRT-based cell division to protein domain architectures.PLoS One. 2022 Mar 31;17(3):e0266395. doi: 10.1371/journal.pone.0266395. eCollection 2022. PLoS One. 2022. PMID: 35358274 Free PMC article.
-
Anchors: A way for FtsZ filaments to stay membrane bound.Mol Microbiol. 2023 Oct;120(4):525-538. doi: 10.1111/mmi.15067. Epub 2023 Apr 28. Mol Microbiol. 2023. PMID: 37503768 Free PMC article. Review.
-
CdrS Is a Global Transcriptional Regulator Influencing Cell Division in Haloferax volcanii.mBio. 2021 Aug 31;12(4):e0141621. doi: 10.1128/mBio.01416-21. Epub 2021 Jul 13. mBio. 2021. PMID: 34253062 Free PMC article.
-
Stable coexistence between an archaeal virus and the dominant methanogen of the human gut.Nat Commun. 2024 Sep 4;15(1):7702. doi: 10.1038/s41467-024-51946-x. Nat Commun. 2024. PMID: 39231967 Free PMC article.
References
-
- Borrel, G., Brugère, J. F., Gribaldo, S., Schmitz, R. A. & Moissl-Eichinger, C. The host-associated archaeome. Nat. Rev. Microbiol. 1–15 10.1038/s41579-020-0407-y (2020). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

