GWAS Mediated Elucidation of Heterosis for Metric Traits in Cotton (Gossypium hirsutum L.) Across Multiple Environments
- PMID: 34093598
- PMCID: PMC8173050
- DOI: 10.3389/fpls.2021.565552
GWAS Mediated Elucidation of Heterosis for Metric Traits in Cotton (Gossypium hirsutum L.) Across Multiple Environments
Abstract
For about a century, plant breeding has widely exploited the heterosis phenomenon-often considered as hybrid vigor-to increase agricultural productivity. The ensuing F1 hybrids can substantially outperform their progenitors due to heterozygous combinations that mitigate deleterious mutations occurring in each genome. However, only fragmented knowledge is available concerning the underlying genes and processes that foster heterosis. Although cotton is among the highly valued crops, its improvement programs that involve the exploitation of heterosis are still limited in terms of significant accomplishments to make it broadly applicable in different agro-ecological zones. Here, F1 hybrids were derived from mating a diverse Upland Cotton germplasm with commercially valuable cultivars in the Line × Tester fashion and evaluated across multiple environments for 10 measurable traits. These traits were dissected into five different heterosis types and specific combining ability (SCA). Subsequent genome-wide predictions along-with association analyses uncovered a set of 298 highly significant key single nucleotide polymorphisms (SNPs)/Quantitative Trait Nucleotides (QTNs) and 271 heterotic Quantitative Trait Nucleotides (hQTNs) related to agronomic and fiber quality traits. The integration of a genome wide association study with RNA-sequence analysis yielded 275 candidate genes in the vicinity of key SNPs/QTNs. Fiber micronaire (MIC) and lint percentage (LP) had the maximum number of associated genes, i.e., each with 45 related to QTNs/hQTNs. A total of 54 putative candidate genes were identified in association with HETEROSIS of quoted traits. The novel players in the heterosis mechanism highlighted in this study may prove to be scientifically and biologically important for cotton biologists, and for those breeders engaged in cotton fiber and yield improvement programs.
Keywords: F1 hybrid; GWAS; hQTNs; heterosis; multiple environments; upland cotton.
Copyright © 2021 Sarfraz, Iqbal, Geng, Iqbal, Nazir, Ahmed, He, Jia, Pan, Sun, Ahmad, Wang, Qin, Liu, Liu, Yang, Ma, Xu, Yang, Zhang, Li, Cai, Zhang, Zhang, Huang, Yi, Zhou, Li, Zhu, Pang, Wang, Sun and Du.
Conflict of interest statement
It is declared that, authors, JL, JY, ZC, and GZ were employed by “Zhongmian Seed Technologies Co., Ltd., Zhengzhou, China,” the author, HL employed by “Jing Hua Seed Industry Technologies Inc., Jingzhou, China,” the author, DX employed by “Guoxin Rural Technical Service Association, Hebei, China,” the authors, JZ as well as LL were employed by “Zhongli Company of Shandong, Shandong, China,” and the author AH employed by “Sanyi Seed Industry of Changde in Hunan Inc., Changde, China.” The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




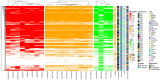

References
-
- Betrán F., Ribaut J., Beck D., De Leon D. G. J. C. S. (2003). Genetic diversity, specific combining ability, and heterosis in tropical maize under stress and nonstress environments. Crop Sci. 43 797–806. 10.2135/cropsci2003.0797 - DOI
-
- Chen G., Du X. (2006). Genetic diversity of basal germplasm phenotypes in upland cotton in China. Acta Bot. Boreali Occident. Sin. 26 1649–1656.
LinkOut - more resources
Full Text Sources

