Adaptive Differences in Gene Expression in Farm-Impacted Seedbeds of the Native Blue Mussel Mytilus chilensis
- PMID: 34093658
- PMCID: PMC8174845
- DOI: 10.3389/fgene.2021.666539
Adaptive Differences in Gene Expression in Farm-Impacted Seedbeds of the Native Blue Mussel Mytilus chilensis
Abstract
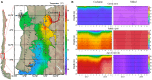
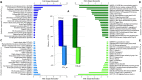
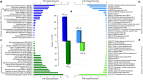
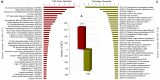
The study of adaptive population differences is relevant for evolutionary biology, as it evidences the power of selective local forces relative to gene flow in maintaining adaptive phenotypes and their underlying genetic determinants. However, human-mediated hybridization through habitat translocations, a common and recurrent aquaculture practice where hybrids could eventually replace local genotypes, risk populations' ability to cope with perturbations. The endemic marine mussel Mytilus chilensis supports a booming farming industry in the inner sea of Chiloé Island, southern Chile, which entirely relies on artificially collected seeds from natural beds that are translocated to ecologically different fattening centers. A matter of concern is how farm-impacted seedbeds will potentially cope with environmental shifts and anthropogenic perturbations. This study provides the first de novo transcriptome of M. chilensis; assembled from tissue samples of mantles and gills of individuals collected in ecologically different farm-impacted seedbeds, Cochamó (41°S) and Yaldad (43°S). Both locations and tissue samples differentially expressed transcripts (DETs) in candidate adaptive genes controlling multiple fitness traits, involved with metabolism, genetic and environmental information processing, and cellular processes. From 189,743 consensus contigs assembled: 1,716 (Bonferroni p value ≤ 0.05) were DETs detected in different tissues of samples from different locations, 210 of them (fold change ≥ | 100|) in the same tissue of samples from a different location, and 665 (fold change ≥ | 4|) regardless of the tissue in samples from a different location. Site-specific DETs in Cochamó (169) and Yaldad (150) in candidate genes controlling tolerance to temperature and salinity shifts, and biomineralization exhibit a high number of nucleotide genetic variants with regular occurrence (frequency > 99%). This novel M. chilensis transcriptome should help assessing and monitoring the impact of translocations in wild and farm-impacted mussel beds in Chiloé Island. At the same time, it would help designing effective managing practices for conservation, and translocation traceability.
Keywords: Mytilus chilensis; adaptative differences; farm-impacted seedbeds; gene expression; genetic variants; transcriptome.
Copyright © 2021 Yévenes, Núñez-Acuña, Gallardo-Escárate and Gajardo.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Epigenetic variation mediated by lncRNAs accounts for adaptive genomic differentiation of the endemic blue mussel Mytiluschilensis.Heliyon. 2023 Dec 14;10(1):e23695. doi: 10.1016/j.heliyon.2023.e23695. eCollection 2024 Jan 15. Heliyon. 2023. PMID: 38205306 Free PMC article.
-
Decoding Local Adaptation in the Exploited Native Marine Mussel Mytilus chilensis: Genomic Evidence from a Reciprocal Transplant Experiment.Int J Mol Sci. 2025 Jan 23;26(3):931. doi: 10.3390/ijms26030931. Int J Mol Sci. 2025. PMID: 39940706 Free PMC article.
-
Adaptive mitochondrial genome functioning in ecologically different farm-impacted natural seedbeds of the endemic blue mussel Mytilus chilensis.Comp Biochem Physiol Part D Genomics Proteomics. 2022 Jun;42:100955. doi: 10.1016/j.cbd.2021.100955. Epub 2022 Jan 4. Comp Biochem Physiol Part D Genomics Proteomics. 2022. PMID: 35065314
-
Mytilus hybridisation and impact on aquaculture: A minireview.Mar Genomics. 2016 Jun;27:3-7. doi: 10.1016/j.margen.2016.04.008. Epub 2016 May 2. Mar Genomics. 2016. PMID: 27157133 Review.
-
Local adaptation and species segregation in two mussel (Mytilus edulis x Mytilus trossulus) hybrid zones.Mol Ecol. 2005 Feb;14(2):381-400. doi: 10.1111/j.1365-294X.2004.02379.x. Mol Ecol. 2005. PMID: 15660932 Review.
Cited by
-
Epigenetic variation mediated by lncRNAs accounts for adaptive genomic differentiation of the endemic blue mussel Mytiluschilensis.Heliyon. 2023 Dec 14;10(1):e23695. doi: 10.1016/j.heliyon.2023.e23695. eCollection 2024 Jan 15. Heliyon. 2023. PMID: 38205306 Free PMC article.
-
Chromosome-Level Genome Assembly of the Blue Mussel Mytilus chilensis Reveals Molecular Signatures Facing the Marine Environment.Genes (Basel). 2023 Apr 7;14(4):876. doi: 10.3390/genes14040876. Genes (Basel). 2023. PMID: 37107634 Free PMC article.
-
Decoding Local Adaptation in the Exploited Native Marine Mussel Mytilus chilensis: Genomic Evidence from a Reciprocal Transplant Experiment.Int J Mol Sci. 2025 Jan 23;26(3):931. doi: 10.3390/ijms26030931. Int J Mol Sci. 2025. PMID: 39940706 Free PMC article.
-
Latitudinal variation and plasticity in response to temperature in Geukensia demissa.Ecol Evol. 2023 Feb 24;13(2):e9856. doi: 10.1002/ece3.9856. eCollection 2023 Feb. Ecol Evol. 2023. PMID: 36844674 Free PMC article.
References
-
- Astorga M., Cárdenas L., Pérez M., Toro J. E., Martínez V., Farías A., et al. (2020). Complex spatial genetic connectivity of mussels Mytilus chilensis along the southeastern pacific coast and Its importance for resource management. J. Shellfish Res. 39 77–86. 10.2983/035.039.0108 - DOI
-
- Astorga M., Vargas J., Valenzuela A., Molinet C., Marín S. L. (2018). Population genetic structure and differential selection in mussel Mytilus chilensis. Aquac. Res. 49 919–927. 10.1111/are.13538 - DOI
LinkOut - more resources
Full Text Sources

